| Issue |
Knowl. Manag. Aquat. Ecosyst.
Number 427, 2026
Anthropogenic impact on freshwater habitats, communities and ecosystem functioning
|
|
|---|---|---|
| Article Number | 14 | |
| Number of page(s) | 11 | |
| DOI | https://doi.org/10.1051/kmae/2026007 | |
| Published online | 10 April 2026 | |
Research Paper
Wastewater treatment effectively eliminates exogenous eDNA, preventing downstream false-positive detections
1
Research Institute for Nature and Forest (INBO), Geraardsbergen, Belgium
2
Ghent University, Department of Biology, Terrestrial Ecology Unit, Ghent, Belgium
3
KU Leuven, Department of Biology, Plant Conservation and Population Biology, Leuven, Belgium
4
Research Institute for Nature and Forest (INBO), Research Center for Aquatic Fauna, Linkebeek, Belgium
* Corresponding author: This email address is being protected from spambots. You need JavaScript enabled to view it.
Received:
28
November
2025
Accepted:
21
March
2026
Abstract
Environmental DNA (eDNA) is widely applied to monitor aquatic biodiversity, but false-positive detections may arise when DNA from consumed or processed organisms is discharged to waterways through domestic wastewater. We experimentally tested whether such wastewater-derived eDNA persists through municipal treatment and contributes to downstream false-positive detections. Controlled spikes of homogenized tissue from six target taxa (two marine fishes and four freshwater fish/amphibians) were introduced into the influent of four wastewater treatment plants (WWTPs) in Flanders, Belgium. Water samples were collected at three points: influent, treated effluent, and the receiving river. Sampling was performed before and up to 24 h after spiking, and eDNA concentrations of each of the six target taxa were quantified using droplet digital PCR. All species showed post-spike increases in influent eDNA concentrations, although individual taxa were not detected in every influent sample, particularly at the largest facilities where hydraulic dilution reduced short-term detectability. In contrast, effluent and downstream concentrations remained zero across all WWTPs, indicating that no spiked eDNA persisted beyond the treatment process. One WWTP located near a marine harbor showed background detection of marine fish species even before spiking, likely originating from seafood processing or domestic discharge. However, these signals were also fully removed during treatment. Overall, modern secondary treatment effectively eliminates exogenous eDNA, preventing its release into receiving waters. While systems with incomplete sewer connectivity or combined sewer overflows may still discharge untreated domestic wastewater, under standard operating conditions riverine eDNA detections can be interpreted as reflecting local biota rather than wastewater contamination.
Key words: Droplet digital PCR (ddPCR) / DNA removal efficiency / combined sewer overflow / environmental monitoring reliability / eDNA transport dynamics
Publisher note: The supplementary material PDF file was corrected on 14 April 2026.
© C. Van Driessche et al., Published by EDP Sciences 2026
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License CC-BY-ND (https://creativecommons.org/licenses/by-nd/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. If you remix, transform, or build upon the material, you may not distribute the modified material.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License CC-BY-ND (https://creativecommons.org/licenses/by-nd/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. If you remix, transform, or build upon the material, you may not distribute the modified material.
1 Introduction
Environmental DNA (eDNA)-based species detection has revolutionized aquatic biodiversity monitoring by enabling the detection of organisms through genetic material emitted into the environment (e.g., released via mucus, scales, feces, gametes, or decaying tissue; Sassoubre et al., 2016) (Thomsen and Willerslev, 2015; Deiner et al., 2017). This approach offers a non-invasive, highly sensitive alternative to traditional monitoring techniques, particularly for assessing fish and amphibian communities across diverse aquatic ecosystems (Hånfling et al., 2016; Valentini et al., 2016). Specifically in lowland rivers and ponds, recent applications have demonstrated the robustness of eDNA for detecting both native and invasive fish and amphibian species under a range of environmental conditions (Brys et al., 2021; Everts et al., 2021; Van Driessche et al., 2022, 2024a).
In lotic environments, eDNA at a given downstream location can originate from multiple upstream inputs, posing challenges for spatially attributing species presence and distinguishing local from transported signals (Jane et al., 2015; Shogren et al., 2017; Van Driessche et al., 2022, 2023). Beyond presence of upstream populations contributing to the eDNA particle pool, eDNA signals may also be supplemented or distorted by a variety of non-local anthropogenic or biological vectors, including trophic transfer, faunal-mediated dispersal, fisheries activities, shipping or ballast-water movement, and broader human-mediated transport processes (Ragot and Villemur, 2022). While the majority of locally detected eDNA originates from resident specimens in the upstream vicinity, hydrological transport and retention processes can carry genetic material over variable distances depending on flow dynamics, substrate type, and degradation rate (Deiner and Altermatt, 2014; Van Driessche et al., 2024a). In addition to these processes, domestic and industrial wastewater constitute consistent and high-volume inputs of exogenous eDNA entering river systems, introducing DNA from consumed species or processed biological materials and thereby creating potential false-positive detections unrelated to the local biota (Inoue et al., 2023; Clare et al., 2022). Recent studies have identified such consumption-derived DNA in wastewater as a credible and underrecognized source of non-local eDNA, capable of persisting through treatment processes and entering receiving water bodies under certain conditions (Xiong et al., 2024b; Inoue et al., 2023). This form of DNA, originating from ingested and excreted tissue, has been shown to contribute to spurious detections in both marine and freshwater systems, especially when using highly sensitive analytical techniques such as qPCR or droplet digital PCR (ddPCR) (Clare et al., 2022; Mauvisseau et al., 2021). These approaches, and particularly multi-marker assays, can further increase detection rates and lower quantification thresholds (Brys et al., 2023), making the distinction between true ecological signals and trace anthropogenic inputs even more critical. This challenge is compounded by the fact that eDNA in wastewater exists in multiple physical states (e.g., dissolved, particle-bound), each with distinct degradation rates influenced by temperature, microbial activity, and water chemistry (Mauvisseau et al., 2021; Barnes et al., 2014). Consequently, interpreting eDNA patterns in urbanized catchments remains complex, as the variable persistence of these different states makes it difficult to distinguish whether detections of common species reflect resident populations or “consumption noise” from domestic discharge. To address this uncertainty, experimental designs must utilize non-local or marine taxa as proxies to explicitly isolate and track the fate of exogenous genetic material across the wastewater-river interface.
Across much of Western Europe, wastewater is treated in wastewater treatment plants (WWTPs) prior to discharge. However, the efficiency of these facilities in removing non-local, perhaps consumption-derived eDNA before it enters natural water bodies remains poorly understood. These facilities receive domestic and industrial wastewater that may carry eDNA from dietary sources or imported species, which may remain detectable if not fully removed during treatment. Modern WWTPs in industrialized regions typically employ a combination of mechanical, biological, and chemical filtration steps designed to remove organic pollutants and microbial loads (Qasim, 2017; Karia and Christian, 2003). Despite these advanced treatments, the persistence of intra- and extracellular eDNA through WWTPs and its potential to confound eDNA-based assessments remains poorly quantified (Nielsen et al., 2007; Caldwell et al., 2021). Although the limited number of studies assessing eDNA in treated effluents has reported variable outcomes, most detections to date appear to be low-level and context-dependent (Inoue et al., 2023; Xiong et al., 2024a, 2024b). Overall, the scarce available evidence suggests that while wastewater-derived eDNA is readily detected in influents, its persistence into effluents is inconsistent and likely influenced by differences in treatment configuration, operational conditions, and analytical sensitivity. Moreover, as wastewater treatment is increasingly regulated to meet standardized performance thresholds in many regions, understanding how effectively such systems prevent the release of non-local eDNA becomes essential, thereby motivating controlled empirical assessments using WWTPs that comply with these standards.
In this study, we aimed to experimentally determine whether consumption-derived eDNA introduced via domestic wastewater can persist through standard WWTP processes and remain detectable in treated downstream water that is discharged in rivers. We conducted a controlled spike experiment at four WWTPs in Flanders, Belgium, encompassing large (Ghent, Ostend) and small (Nevele, Zomergem) facilities with varying treatment capacities but using the same procedure to remove organic pollutants and microbial loads. A predefined mixture of species, comprising commonly consumed flatfish (Solea solea and Pleuronectes platessa), a native but rare freshwater fish species (Misgurnus fossilis), and three non-native species even more unlikely to occur in the environment (i.e., Misgurnus bipartitus, Xenopus laevis, and Lithobates catesbeianus), was introduced into the influent stream of each facility. Water samples for eDNA analyses were subsequently taken before and after spiking at three points: the influent, the treated effluent, and the receiving river 100 meters downstream of the WWTP effluent. We then used ddPCR, a highly sensitive molecular technique suitable for quantifying species-specific trace DNA levels (Doi et al., 2015; Hindson et al., 2011). We hypothesized the WWTP treatment chain effectively removes consumption-derived eDNA from wastewater. By explicitly evaluating the detectability and spatial persistence of experimentally introduced and highly concentrated target DNA, this work offers new empirical insights into the accuracy and reliability of eDNA-based monitoring in riverine and lowland aquatic ecosystems. It also offers practical insights for interpreting eDNA data in human-impacted catchments, with potential relevance for eDNA surveillance protocols, water quality assessments, and environmental regulation.
2 Materials and methods
2.1 Experimental set-up
This study was performed on the 9th and 10th of September, 2024, at four WWTPs located in Flanders, Belgium: Ostend, Ghent, Nevele, and Zomergem (Fig. 1). Although the filtering procedure steps are the same, the selected WWTPs differ markedly in terms of capacity and population equivalents served. Ostend and Ghent represent large facilities, treating 50 099 m3/day and 78 720 m3/day during the experiment, respectively, corresponding to 198 000 and 207 000 population equivalents. The two smaller facilities at Nevele and Zomergem, on the other hand, treat 4351 m3/day and 1877 m3/day, with respective population equivalents of 9000 and 5400. At each WWTP, eDNA water samples were collected from three site-specific locations: (i) the influent (i.e., incoming untreated wastewater), (ii) the effluent (i.e., treated water discharged by the facility), and (iii) a site in the river 100 m downstream of the WWTP effluent (Fig. 1).
At the onset of the experiment, eDNA samples were taken at each of these three site-specific locations at every WWTP. For this eDNA-sampling, water was collected from the influent and the effluent using the facilities’ automated collectors, which provide a standardized 24-h composite sample representing the full daily wastewater cycle. For the sampling in the river downstream of the site, water was collected cross-sectionally to the river using a 0.5-L sterile sampling bag attached to a telescopic sampling pole. To capture the expected passage of the spiked material through the treatment system, 15–20 subsamples were collected in the river downstream of the site at 12 h and again at 24 h post-spiking, corresponding to the typical hydraulic retention time (HRT) of medium-scale activated-sludge WWTPs, which generally ranges from ∼8 to 24 h depending on plant size and flow conditions (Qasim, 2017; Aquafin, pers. comm.). All subsamples were combined and homogenized to form a pooled water sample for each location. From each sample, 2 L of water was then filtered through an enclosed disk filter containing an integrated 5 μm glass fiber prefilter and a 0.8 μm PES membrane (NatureMetrics, Surrey, England). At the end of each filtration, the remaining water inside the capsule was expelled by pushing air through the filter capsule until it dried completely. The filters were then capped at both ends and stored at −21∘C until further laboratory processing.
Next, a controlled eDNA “spike” was introduced into the influent stream of each WWTP. This spike (Fig. 2) consisted of a homogenized tissue mixture comprising six species: two commonly consumed flatfish species (Solea solea and Pleuronectes platessa) and four non-consumed species, including the native Misgurnus fossilis and the non-native invasive Misgurnus bipartitus, Xenopus laevis, and Lithobates catesbeianus. Those non-native taxa were deliberately chosen because they do not occur naturally in the system, thereby enabling clear discrimination between experimentally introduced DNA and any background eDNA originating from genuine wastewater inputs. In contrast, the two flatfish species were included because they represent realistic, consumption-derived taxa that may occur in domestic wastewater, thereby reflecting plausible real-world sources of exogenous eDNA entering WWTPs. The flatfish specimens were sourced from a local fish market, while M. fossilis was obtained from the conservation breeding facility at the INBO Research Center for Aquatic Fauna (Linkebeek). Specifically, these M. fossilis specimens consisted of naturally deceased individuals that originated from routine cultivation and harvesting operations in the breeding facility’s rearing ponds (Van Wichelen et al., 2023). The invasive M. bipartitus, X. laevis, and L. catesbeianus were captured specimens retrieved through an invasive species management program. No animals were killed for the specific purpose of this experiment but obtained as already deceased individuals. All specimens were blended to produce a uniform mixture, which was then equally divided into four equal portions, one for each WWTP. Because this spike consisted of raw homogenized tissue rather than digested material, it contained far higher DNA quantities than would typically be present in domestic wastewater, where consumption-derived eDNA is strongly degraded and diluted. This intentional use of a high-yield source material was designed to represent a conservative, worst-case scenario for testing removal efficiency. Once introduced into the influent stream, however, the spike was immediately dispersed into very high flow volumes, meaning that the effective concentration entering the treatment system was strongly reduced. Given the high flow rates and turbulent mixing at influent inlets, particularly in large facilities, the spike likely behaved as a short-lived pulse rather than an instantaneously homogenized solution, further diluting the introduced material within the 24−h composite influent samples.
Twenty-four hours after this spiking event (T2, Fig. 1), water samples were again collected at the same site-specific locations per WWTP and filtered for collection of eDNA following the same procedure as described for the pre-spiking samples. In total, 24 eDNA samples were collected.
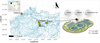 |
Fig. 1 Overview of the wastewater treatment plant (WWTP) experiment. eDNA samples were collected at four WWTPs in Belgium (Ostend, Ghent, Zomergem, and Nevele) at three sampling locations per WWTP: the influent (untreated wastewater), the effluent (treated water leaving the WWTP), and the receiving river 100 m downstream. A predefined DNA spike was introduced at the influent of each WWTP, with samples collected before (T1) and after (T2) the spiking event to assess the persistence and detectability of consumption-derived fish DNA following wastewater treatment. The map shows the location of the four WWTPs within Flanders, with major waterways indicated in blue. The schematic on the right illustrates the sampling strategy within the WWTP system. Base illustration of the wastewater treatment plant adapted from NoBrokerHood (2021). |
 |
Fig. 2 Composition of the eDNA spike introduced at the influent of each WWTP. Each spike contained a predefined mixture of six species and their biomasses, including two commonly consumed flatfish species (Solea solea and Pleuronectes platessa) as well as the rare Misgurnus fossilis and three non-native species (i.e., Misgurnus bipartitus, Xenopus laevis, and Lithobates catesbeianus) that are unlikely to occur naturally in the river or to be present in human diets. |
2.2 Laboratory procedures
DNA extraction from the filters was conducted according to Brys et al. (2021), using the SX capsule method with the DNeasy Blood & Tissue Kit (Qiagen), following the manufacturer’s guidelines. This extraction process was performed in a dedicated PCR-free room for low-copy-number template extractions, with particulate air-filtered compartments, avoiding contamination of eDNA samples. The resulting eDNA extract was quantified prior to amplification with a Quantus Fluorometer according to the manufacturer’s instructions.
Species-specific eDNA detection and quantification of each of the six target species were obtained using ddPCR on a QX200 ddPCR system (Bio-Rad, Temse, Belgium) in 20 μL reaction volumes. The species-specific primer/probe assays used in this study were previously developed and validated, and are further detailed in Table 1. Final eDNA concentrations per assay were obtained as described in (Brys et al., 2021; Everts et al., 2021; Van Driessche et al., 2022). For each of the six target species assays, all 24 water samples were analyzed in triplicate, resulting in three technical replicates. Each ddPCR plate included extraction blanks, no-template controls, and an assay-specific positive control consisting of diluted tissue-derived reference DNA. eDNA concentrations (copies μL−1) were calculated using Poisson correction of positive droplet counts (QuantaSoft).
To provide broader context on the background fish community and validate ddPCR results, parallel eDNA metabarcoding was conducted on all samples using 12S rRNA primers (see Supplementary Information S1 for details).
Characteristics of the primer/probe assays used for the detection of Solea solea, Pleuronectes platessa, Misgurnus fossilis, Misgurnus bipartitus, Xenopus laevis, and Lithobates catesbeianus, respectively.
2.3 Statistical analyses
All statistical analyses were conducted in R version 4.5.1 (R Core Team, 2025) using the packages Ime4 v1.1.37 (Bates et al., 2015), ImerTest v3.1.3 (Kuznetsova et al., 2017), emmeans v1.11.2.8 (Lenth, 2023), and dplyr v1.1.4 (Wickham, 2023). Analyses focused on evaluating whether eDNA introduced during the spike event was effectively removed by wastewater treatment and whether any residual DNA could be detected in downstream waters. Figures were created using the additional package ggplot2 v3.5.2 (Wickham, 2016).
Because effluent and downstream samples produced no detections, the presence–absence analysis was restricted to the influent. Pre- and post-spike detection frequencies were compared using a Fisher’s exact test. Secondly, the quantitative analysis used ddPCR concentration data from all locations, as zero concentrations in effluent and downstream samples represent meaningful values for evaluating treatment performance. To evaluate quantitative variation in eDNA concentration, a linear mixed-effects model (LMM) was fitted to the log-transformed ddPCR concentrations. eDNA concentrations (copies μL−1) were transformed using log10(x+1) to improve normality and reduce the influence of zero concentrations. The fixed-effects structure included sampling location (influent, effluent, and downstream), sampling period (pre- or post-spiking), and their interaction, whereas WWTP identity was included as a random intercept to account for repeated measurements within facilities:
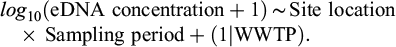 (1)
(1)
Model residuals were visually inspected to verify approximate normality and homoscedasticity. Estimated marginal means (EMMs) and Tukey-adjusted pairwise contrasts were obtained using the emmeans package to compare differences in mean eDNA concentration among locations and sampling periods.
3 Results
3.1 Data quality
No amplification was observed in extraction blanks or PCR negative controls for any of the spiked target taxa, indicating the absence of detectable contamination during field sampling, DNA extraction, and amplification procedures.
3.2 Detection patterns and eDNA removal
Across all four WWTPs, background eDNA levels of the spiked taxa were largely absent prior to the experiment (pre-spiking), confirming minimal existing contamination prior to the onset of the experiment. An exception occurred at the Ostend WWTP, where low eDNA concentrations of Solea solea and Pleuronectes platessa were detected in the pre-spike influent samples (Figs. 3 and 4). These detections likely originated from local seafood processing or domestic discharge, consistent with the city’s proximity to a major fishing harbor (Ostend Fish Market, Port of Ostend). Such background detections illustrate the relevance of the research question and the realistic presence of consumption-derived marine eDNA in untreated wastewater.
Across all facilities, influent samples showed more detections after the spike than before (Fig. 3). A Fisher’s exact test confirmed a significant increase in influent detection probability after the spike (odds ratio = 6.54, 95% CI = 2.52−19.31, P < 0.0001). Effluent and downstream samples showed no detections at any facility or sampling time.
At the two smaller facilities (Nevele and Zomergem), all taxa were consistently detected, whereas the larger WWTPs (Ghent and Ostend) showed occasional non-detections (Fig. 4). Log-transformed eDNA concentrations (log10[copies μ−1 + 1]) showed a clear and location-specific response to the spike event. The linear mixed-effects model showed significant effects of sampling location (F = 40.56, P < 0.001), spike period (F = 12.49, P < 0.001), and their interaction (F = 12.49, P < 0.001). Estimated marginal means indicated that influent concentrations increased after the spike (pre: 0.094 ± 0.034; post: 0.329 ± 0.034), while effluent and downstream concentrations remained near zero at both sampling periods (Fig. 4). Tukey-adjusted pairwise contrasts showed that the spike significantly increased eDNA concentrations only in the influent (P < 0.0001). Concentrations did not differ between effluent and downstream sites (all P > 0.05).
Metabarcoding results corroborated these ddPCR findings (Supplementary Fig. S1). While resident fish communities were successfully characterized across all sites, the experimentally introduced target species were only detectable via eDNA metabarcoding in the Ostend site pre-spiking and in the influent samples across all WWTP locations post-spiking.
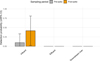 |
Fig. 3 Overall treatment barrier effect: Mean eDNA detection probability (±95% CI) of the six spiked species across WWTPs, based on the proportion of positive ddPCR detections before (pre-spike) and after (post-spike) the controlled introduction of the homogenized tissue mixture at the influent. Bars show mean detection probabilities aggregated across all taxa and facilities. The X-axis represents the sampling locations per WWTP, i.e., the influent (incoming untreated wastewater), the effluent (treated discharge water), and the downstream river site (100 m from the effluent outflow). |
 |
Fig. 4 Measured eDNA concentrations (copies per μL) of the six spiked species (i.e., Xenopus laevis, Misgurnus fossilis, Misgurnus bipartitus, Pleuronectes platessa, Lithobates catesbeianus, and Solea solea) across four WWTPs (Ghent, Ostend, Nevele, and Zomergem) at three sampling locations per WWTP: influent (incoming wastewater), effluent (treated water released from the WWTP), and downstream river (100 m from effluent discharge point). The shaded areas with blender icons indicate the introduction of the DNA spike containing six species. Box plots show the distribution of eDNA concentrations for each species across filter replicates. Note that the Y-axis range differs for each panel. |
4 Discussion
This study provides clear experimental evidence that consumption-derived environmental DNA (eDNA), introduced at relatively high concentrations via domestic or household wastewater, can effectively be removed during standard secondary wastewater treatment. Across all four WWTPs, eDNA from the spiked species showed increased concentrations in the post-spike influent samples, although detections were not uniformly present for every species at every facility. In both effluent and downstream samples, however, all species were consistently undetectable. Detection frequencies increased significantly in the influent after the spike, as confirmed by the Fisher’s exact test, while no detections were observed in either effluent or downstream samples. Quantitatively, influent eDNA concentrations were significantly higher than those measured at subsequent treatment stages, as shown by the linear mixed-effects model and Tukey contrasts. Together, these findings demonstrate that modern WWTP processes are highly efficient at removing exogenous DNA, thereby minimizing the potential for false-positive detections of non-local fish or amphibian species in downstream waters.
Interestingly, background detections at the Ostend facility, where marine fish species such as Solea solea and Pleuronectes platessa were identified in pre-spike influent samples, highlight the realistic presence of consumption-derived or industrially sourced eDNA in urban wastewater streams. These detections likely originate from seafood processing or domestic discharge associated with Ostend’s coastal economy. Similar observations have been reported in other urban and coastal settings, where DNA from market fish, aquaculture products, or industrial effluents was detected in influent or sewage samples (Inoue et al., 2023; Clare et al., 2022). Comparable coastal cases have also been described by Van Driessche et al. (2024b), where eDNA from marine and estuarine species reflected local economic and hydrological linkages rather than ecological presence. Importantly, no corresponding signals were observed in effluent or receiving waters, confirming that the treatment process acts as an effective barrier against discharge of exogenous DNA. This aligns with findings from Clare et al. (2022) and Zhang et al. (2025).
Parallel metabarcoding (Supplementary Fig. S1) detected the target spike species in post-spike influent samples, though relative read abundance remained low compared to dominant local species, and there was no targeted detection in effluent or downstream river samples. This illustrates that while metabarcoding is effective for broad biodiversity assessment, targeted ddPCR is more robust for quantification of trace-level DNA in complex matrices. As noted by Wood et al. (2019), targeted assays are less influenced by primer competition or taxonomic bias in high-biomass environments. Together, these results highlight the robustness of ddPCR and the reliability of eDNA-based monitoring in anthropogenically influenced systems. Furthermore, even if trace eDNA particles bypassed treatment, its ecological impact would likely remain negligible. In lotic systems, locally emitted and less fragmented resident biota typically dominate eDNA patterns. These local signals tend to overwhelm distant or degraded anthropogenic inputs in relative read abundance data (Van Driessche et al., 2023). This reinforces the reliability of eDNA monitoring, as well-functioning infrastructure prevents exogenous DNA from distorting the local ecological signal.
In our experiment, post-dilution influent concentrations ranged from approximately 3 to 60 copies μL−1 (Fig. 4), equivalent to 3 × 106 to 6 × 107 copies L−1. These values fall within, or even exceed, the upper ranges reported for fish mitochondrial eDNA in wastewater-associated environments (Inoue et al., 2023). Because our spike consisted of undigested homogenized tissue, it represents a conservative, high-load scenario compared to genuine consumption-derived eDNA, which undergoes significant degradation during digestion and sewer transport. The complete absence of detectable eDNA in effluent and receiving waters despite these high influent loads indicates that standard secondary treatment effectively retains and degrades exogenous DNA before discharge. This high removal efficiency is driven by the microbial purification inherent to the activated sludge process. Our results align with Xiong et al. (2024a), who demonstrated that the majority of extracellular DNA is removed through sorption to extracellular polymeric substances (EPS) and subsequent microbial degradation within the biological treatment stage. Mechanistically, this process acts as a critical filter where DNA is removed via a combination of physical retention in primary sedimentation, sorption to activated sludge flocs, and enzymatic cleavage during aeration (Nielsen et al., 2007; Caldwell et al., 2021; Xiong et al., 2024a). These findings mirror eDNA decay mechanisms observed in natural sedimentary environments (Jo et al., 2023). While the homogenized tissue used in this study primarily contained DNA protected within cellular or organellar membranes, the complete removal observed suggests that dissolved DNA, which lacks such protection and is more susceptible to enzymatic cleavage, would be eliminated even more rapidly. Our experiment thus represents a conservative assessment, as intracellular DNA is typically more stable than the dissolved fragments often associated with highly degraded anthropogenic inputs (Xiong et al., 2024a; Nielsen et al., 2007). Recent synthesis work identifies anthropogenic eDNA inputs, including wastewater and urban runoff, as an important challenge for obtaining accurate and ecologically interpretable molecular biodiversity data (Çevik and Çevik, 2025). Our findings add nuance to this ongoing discussion about wastewater as a potential source of non-local DNA that could confound eDNA monitoring surveys in adjacent rivers. By demonstrating that even elevated influent concentrations are fully removed, our results show that well-functioning secondary treatment largely mitigates this source of uncertainty.
Regional and infrastructural differences are likely to influence the extent to which wastewater contributes detectable eDNA to the aquatic environment. In our study, these infrastructural effects manifested clearly at the influent stage of the larger facilities, where the spike was diluted beyond full species detectability. This reduced detectability at large facilities is consistent with the hydraulic behavior of influent streams, where very high flow rates and turbulence across upstream inlet structures (e.g., screw pumps, grit chambers, and inlet channels) prevent instantaneous homogenization of the added material (Metcalf and Eddy, 2014; Tchobanoglous Leverenz, 2013). Under such conditions, a spike behaves as a short-lived pulse rather than a uniformly mixed signal, a dynamic well documented for shock or slug loads in large facilities (Metcalf and Eddy, 2014). Consequently, the introduced spike can be temporarily unevenly dispersed or strongly diluted within the 24-h composite samples, which are known to dampen short-term concentration peaks (Ort et al., 2010). While this ‘averaging’ effect dampens the measured concentration of a short-lived pulse, it provides a quantitatively robust measure of the total DNA mass load handled by the facility. By capturing the integrated mass of the spike in the influent, which remained significantly elevated despite dilution, and observing its total absence in the effluent, we can confirm a complete removal efficiency. This mass-balance approach ensures that our conclusions reflect the plant’s overall processing capacity rather than transient peak dynamics. Such mixing-related variability is therefore an important consideration when interpreting influent eDNA concentrations but does not compromise assessments of downstream treatment performance. In regions with lower treatment efficiency, incomplete sewer connectivity, or frequent combined sewer overflows, influent concentrations of consumption-derived DNA may be higher, and a greater proportion of untreated material may enter natural waterways (Inoue et al., 2023; Xiong et al., 2024b). Such overflows occur when heavy rainfall or high hydraulic loads exceed treatment capacity, causing temporary bypasses that discharge untreated domestic and stormwater directly into rivers (Mailhot et al., 2015). These episodic events can release dietary or industrially derived eDNA into receiving waters, potentially leading to short-lived, spatially restricted false-positive detections. Similarly, households not connected to centralized WWTP networks, which is still common in rural or developing areas, may discharge greywater directly into streams, potentially introducing dietary or domestic eDNA (Khajvand et al., 2022; Sha et al., 2024). Such variability underscores the importance of local infrastructure and hydrological context when interpreting eDNA detections near populated areas. For ecological surveys, sampling designs should therefore account for proximity to wastewater outfalls and hydrological connectivity. Collecting samples either upstream or sufficiently downstream from effluent discharge zones, or verifying flow separation from urban drainage, can help ensure that eDNA signals are representative of local biota rather than anthropogenic inputs. Comparative studies across regions differing in treatment technologies, discharge regulation, and catchment type will be essential to generalize these findings and refine eDNA-based monitoring frameworks under varying environmental conditions.
Overall, our results show that secondary wastewater treatment effectively eliminates consumption-derived DNA, preventing its release into receiving waters. Consequently, effluents from modern WWTPs are unlikely to generate false-positive eDNA detections downstream, reinforcing the reliability of eDNA-based biodiversity monitoring in urbanized landscapes. These conclusions apply to well-functioning treatment systems in lowland temperate regions. Areas with higher wastewater loads, intensive seafood processing, or incomplete sewer networks may exhibit different dynamics and warrant further study. By demonstrating complete removal of exogenous DNA under realistic operational conditions, this work strengthens confidence in interpreting riverine eDNA signals as reflections of local biota rather than wastewater contamination. Future efforts should integrate molecular and hydrological approaches across facilities of varying design to refine predictions of eDNA persistence and transport under diverse environmental conditions.
Acknowledgments
We gratefully acknowledge Aquafin NV and, in particular, Sarah Govaert for their valuable collaboration and logistical support throughout this study, including access to wastewater treatment facilities and assistance with sampling coordination. We also thank the on-site technical staff for facilitating the experimental setup and ensuring safe and efficient operations during fieldwork.
Funding
This work was supported by Research Foundation Flanders (FWO) [No. 1S23822N and No. 1S01822N].
Conflicts of interest
The authors declare that they have no conflicts of interest in relation to this article.
Data availability statement
All data are available from the Zenodo Digital Repository (https://doi.org/10.5281/zenodo.17952416).
Author contribution statement
Charlotte Van Driessche: Conceptualization, Methodology, Software, Formal analysis, Investigation, Writing – Original Draft, Visualization, Supervision, Project administration, Funding acquisition. Teun Everts: Investigation, Writing – Review & Editing, Funding Acquisition. Berdien De Beer: Investigation, Writing – Review & Editing. Sabrina Neyrinck: Investigation, Writing – Review & Editing. Johan Auwerx: Investigation, Writing – Review & Editing. Dries Bonte: Validation, Supervision, Writing – Review & Editing. Rein Brys: Conceptualization, Investigation, Supervision, Writing – Review & Editing.
Supplementary Material
Figure S1. Spatiotemporal fish community composition across four wastewater treatment plants (WWTPs) based on eDNA metabarcoding. Relative read abundance is shown for three sampling points per WWTP (influent, effluent, and downstream river), both before and after a controlled spike of six target species. Red stars indicated the detection of at least one of the six target species (Misgurnus fossilis, Solea solea, Pleuronectes platessa, Xenopus laevis, and Lithobates catesbeianus). Note that target species were detected across all influent post-spike samples but were absent from all effluent and downstream river samples. Access Supplementary Material
References
- Barnes MA, Turner CR, Jerde CL, Renshaw MA, Chadderton WL, Lodge DM. 2014. Environmental conditions influence eDNA persistence in aquatic systems. Environ Sci Technol 48: 1819–1827. [Google Scholar]
- Bates D, Måchler M, Bolker B, Walker S. 2015. Fitting linear mixed-effects models using Ime4. J Stat Softw 67: 1–48. [CrossRef] [Google Scholar]
- Brys R, Haegeman A, Halfmaerten D, Neyrinck S, Staelens A, Auwerx J, Ruttink T. 2021. Monitoring of spatiotemporal occupancy patterns of fish and amphibian species in a lentic aquatic system using environmental DNA. Mol Ecol 30: 3097–3110. [Google Scholar]
- Brys R, Halfmaerten D, Everts T, Van Driessche C, Neyrinck S. 2023. Combining multiple markers significantly increases the sensitivity and precision of eDNA-based single-species analyses. Environ DNA, 5: 1065–1077. [Google Scholar]
- Caldwell JM, Raley ME, Levine BR. 2021. Detection and quantification of DNA from wastewater: A new tool for understanding human-wildlife interactions. Environ Sci Technol 55: 1244–1252. [Google Scholar]
- Çevik T, Çevik N. 2025. Environmental DNA (eDNA): a review of ecosystem biodiversity detection and applications. Biodivers Conserv 34: 2999–3035. [Google Scholar]
- Clare FC, Economou CK, Faulkes Z, Seymour M, Creer S. 2022. Wastewater as a potential source of eDNA contamination in aquatic monitoring: considerations from consumption-derived DNA. Environ DNA 4: 23–34. [Google Scholar]
- Deiner K, Altermatt F. 2014. Transport distance of invertebrate environmental DNA in a natural river. PLOS One 9: e88786. [Google Scholar]
- Deiner K, Bik HM, Måchler E, Seymour M, Lacoursière-Roussel A, Altermatt F, Creer S, Bista I, Lodge DM, de Vere N, Pfrender ME, Bernatchez, L. 2017. Environmental DNA metabarcoding: transforming how we survey animal and plant communities. Mol Ecol 26: 5872–5895. [CrossRef] [PubMed] [Google Scholar]
- Doi H, Uchii K, Takahara T, Matsuhashi S, Yamanaka H, Minamoto T. 2015. Use of droplet digital PCR for estimation of fish abundance and biomass in environmental DNA surveys. PLOS One 10: e0122763. [Google Scholar]
- Everts T, Halfmaerten D, Neyrinck S, De Regge N, Jacquemyn H, Brys R. 2021. Accurate detection and quantification of seasonal abundance of American bullfrog (Lithobates catesbeianus) using ddPCR eDNA assays. Sci Rep 11: 11282. [Google Scholar]
- Hånfling B, Handley LL, Read DS, Hahn C, Li J, Nichols P, Blackman RC, Oliver A, Winfield IJ. 2016. Environmental DNA metabarcoding of lake fish communities reflects long-term data from established survey methods. Mol Ecol 25: 3101–3119. [Google Scholar]
- Hindson BJ, Ness KD, Masquelier DA, Belgrader P, Heredia NJ, Makarewicz A et al. 2011. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem 83: 8604–8610. [Google Scholar]
- Inoue D, Ushio M, Murakami H, Miya M. 2023. Consumption-derived eDNA in human sewage as a potential source of contamination in aquatic biodiversity monitoring. Sci Total Environ 858: 159770. [Google Scholar]
- Jane SF, Wilcox TM, McKelvey KS, Young MK, Schwartz MK, Lowe WH, Letcher BH, Whiteley AR. 2015. Distance, flow, and PCR inhibition: eDNA dynamics in two headwater streams. Mol Ecol Resour 15: 216–227. [Google Scholar]
- Jo TS. 2023. Utilizing the state of environmental DNA (eDNA) to incorporate time-scale information into eDNA analysis. Proc Royal Soc B 290: 20230979. [Google Scholar]
- Karia GL, Christian RA. 2003. Wastewater Treatment: Concepts and Design Approach. Prentice-Hall of India. [Google Scholar]
- Khajvand M, Mostafazadeh AK, Drogui P, Tyagi RD. 2022. Management of greywater: environmental impact, treatment, resource recovery, water recycling, and decentralization. Water Sci Technol 86: 909–937. [Google Scholar]
- Knudsen SW, Ebert RB, Hesselsøe M, Kuntke F, Hassingboe J, Mortensen PB, et al. 2019. Species-specific detection and quantification of environmental DNA from marine fishes in the Baltic Sea. J Exp Mar Biol Ecol 510: 31–45. [Google Scholar]
- Kuznetsova A, Brockhoff PB, Christensen RH. 2017. ImerTest package: tests in linear mixed effects models. J Stat Softw 82: 1–26. [Google Scholar]
- Lenth R. 2023. emmeans: Estimated Marginal Means, aka Least-Squares Means_. R package version 1.8. 5. [Google Scholar]
- Lin M, Zhang S, Yao M. 2019. Effective detection of environmental DNA from the invasive American bullfrog. Biol Invasions 21: 2255–2268. [Google Scholar]
- Maes SM, Desmet S, Brys R, Sys K, Ruttink T, Maes S, Hostens K, Vansteenbrugge L, Derycke, S. 2024. Detection and quantification of two commercial flatfishes (Solea solea and Pleuronectes platessa) in the North Sea using environmental DNA. Environ DNA 6: e426. [Google Scholar]
- Mailhot A, Talbot G, Lavallée B. 2015. Relationships between rainfall and Combined Sewer Overflow (CSO) occurrences. J Hydrol 523: 602–609. [Google Scholar]
- Mauvisseau Q, Halfmaerten D, Neyrinck S, Burian A, Brys R. 2021. Effects of preservation strategies on environmental DNA detection and quantification using ddPCR. Environmental DNA, 3, 815–822. [Google Scholar]
- Metcalf & Eddy. 2014. Wastewater engineering: treatment and resource recovery, McGrawHill Education, New York, 2018 p. [Google Scholar]
- Nielsen KM, Johnsen PJ, Bensasson D, Daffonchio D. 2007. Release and persistence of extracellular DNA in the environment. Environ Biosafety Res 6: 37–53. [Google Scholar]
- NoBrokerHood. 2021. Sewage Treatment Plant – What It Is and How It Works. Retrieved from https://www.nobrokerhood.com/blog/sewage-treatment-plant/ [Google Scholar]
- Ort C, Lawrence MG, Reungoat J, Mueller JF. 2010. Sampling for PPCPs in wastewater systems: comparison of different sampling modes and optimization strategies. Environ Sci Technol 44: 6289–6296. [Google Scholar]
- Qasim SR. 2017. Wastewater Treatment Plants: Planning, Design, and Operation. Routledge. [Google Scholar]
- R Core Team (2025). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. [Google Scholar]
- Ragot SA, Villemur R. 2022. Potential for non-local eDNA inputs through predator dispersal in aquatic ecosystems. Front Environ Sci 10: 891005. [Google Scholar]
- Sassoubre LM, Yamahara KM, Gardner LD, Block BA, Boehm AB. 2016. Quantification of environmental DNA (eDNA) shedding and decay rates for three marine fish. Environ Sci Technol 50: 10456–10464. [Google Scholar]
- Secondi J, Dejean T, Valentini A, Audebaud B, Miaud C. 2016. Detection of a global aquatic invasive amphibian, Xenopus laevis, using environmental DNA. Amphib-Rept 37: 131–136. [Google Scholar]
- Sha C, Shen S, Zhang J, Zhou C, Lu X, Zhang H. 2024. A review of strategies and technologies for sustainable decentralized wastewater treatment. Water 16: 3003. [Google Scholar]
- Shogren AJ, Tank JL, Egan SP, August OM, Rosi EJ. 2017. Controls on eDNA movement in streams: Transport, retention, and resuspension. Sci Rep 7: 5065. [Google Scholar]
- Strickler KM, Fremier AK, Goldberg CS. 2015. Quantifying effects of UV-B, temperature, and pH on eDNA degradation in aquatic microcosms. Biol Conserv 183: 85–92. [Google Scholar]
- Tchobanoglous G, Leverenz H. 2013. The rationale for decentralization of wastewater infrastructure. In: T.A. Larsen, K.M. Udert, & J. Lienert (Eds.), Separation and Decentralization for Wastewater Management, IWA Publishing, London, UK. pp. 101–115. [Google Scholar]
- Thomsen PF, Willerslev E. 2015. Environmental DNA – an emerging tool in conservation for monitoring past and present biodiversity. Biol Conserv 183: 4–18. [Google Scholar]
- Valentini A, Taberlet P, Miaud C, et al. 2016. Next-generation monitoring of aquatic biodiversity using environmental DNA metabarcoding. Mol Ecol 25: 929–942. [CrossRef] [MathSciNet] [PubMed] [Google Scholar]
- Van Driessche C, Everts T, Neyrinck S, Brys R. 2022. Experimental assessment of downstream environmental DNA patterns under variable fish biomass and river discharge rates. Environ DNA 169: 112807. [Google Scholar]
- Van Driessche C, Everts T, Neyrinck S, Deflem I, Bonte D, Brys R. 2024a. Reduced sampling intensity through key sampling site selection for optimal characterization of riverine fish communities by eDNA metabarcoding. Ecol Indic 169: 112807. [Google Scholar]
- Van Driessche C, Everts T, Neyrinck S, Halfmaerten D, Haegeman A, Ruttink T, Bonte D, Brys R. 2023. Using environmental DNA metabarcoding to monitor fish communities in small rivers and large brooks: insights on the spatial scale of information. Environ Res 228: 115857. [Google Scholar]
- Van Driessche C, Everts T, Neyrinck S, Halfmaerten D, Verschelde P, Breine J, Bonte D, Brys R. 2024b. Environmental DNA metabarcoding reflects spatiotemporal fish community shifts in the Scheldt estuary. Sci Total Environ 934: 173242. [Google Scholar]
- Van Wichelen J, Auwerx J, Maes Y, Brys R, Mergeay J, Maes D, De Bruyn L, Verreycken H, Belpaire C, de Bruin A, Vos, M. 2023. Achtergrondrapport van het soortenbeschermingsprogramma voor grote modderkruiper (Misgurnus fossilis Linnaeus 1758) in Vlaanderen (2021–2025). [Google Scholar]
- Wickham H, François R, Henry L, Müller K, Vaughan D. 2023. dplyr: A Grammar of Data Manipulation. https://doi.org/10.32614/CRAN.package.dplyr [Google Scholar]
- Wickham H, Chang W, Wickham MH. 2016. Package ‘ggplot2’. Create elegant data visualisations using the grammar of graphics. Version 2, 1–189. [Google Scholar]
- Wood SA, Pochon X, Laroche O, von Ammon U, Adamson J, Zaiko A. 2019. A comparison of droplet digital polymerase chain reaction (PCR), quantitative PCR and metabarcoding for species-specific detection in environmental DNA. Mol Ecol Resour 19: 1407–1419. [Google Scholar]
- Xiong W, Maclsaac HJ, Zhan A. 2024a. An overlooked source of false positives in eDNA-based biodiversity assessment and management. J Environ Manag 358: 120949. [Google Scholar]
- Xiong W, Jin Y, Wang Y, Wang S, Chen B, Su H. 2024b. Novel insights into the biological state in algal-bacterial granular sludge granulation: armor-like protection provided by the algal barrier. Water Res 262: 122087. [Google Scholar]
- Zhang S, Al-Gashgari B, Medina JS, Wang C, Narayanasamy S, Xiong Y, Wang K, Hong PY. 2025. Extracellular DNA-associated dissemination of antimicrobial resistance in anaerobic versus aerobic membrane bioreactor. Bioresour Technol 437: 133054. [Google Scholar]
Cite this article as: Van Driessche C, Everts T, De Beer B, Neyrinck S, Auwerx J, Bonte D, Brys R. 2026. Wastewater treatment effectively eliminates exogenous eDNA, preventing downstream false-positive detections. Knowl. Manag. Aquat. Ecosyst., 427, 14. https://doi.org/10.1051/kmae/2026007
All Tables
Characteristics of the primer/probe assays used for the detection of Solea solea, Pleuronectes platessa, Misgurnus fossilis, Misgurnus bipartitus, Xenopus laevis, and Lithobates catesbeianus, respectively.
All Figures
 |
Fig. 1 Overview of the wastewater treatment plant (WWTP) experiment. eDNA samples were collected at four WWTPs in Belgium (Ostend, Ghent, Zomergem, and Nevele) at three sampling locations per WWTP: the influent (untreated wastewater), the effluent (treated water leaving the WWTP), and the receiving river 100 m downstream. A predefined DNA spike was introduced at the influent of each WWTP, with samples collected before (T1) and after (T2) the spiking event to assess the persistence and detectability of consumption-derived fish DNA following wastewater treatment. The map shows the location of the four WWTPs within Flanders, with major waterways indicated in blue. The schematic on the right illustrates the sampling strategy within the WWTP system. Base illustration of the wastewater treatment plant adapted from NoBrokerHood (2021). |
| In the text | |
 |
Fig. 2 Composition of the eDNA spike introduced at the influent of each WWTP. Each spike contained a predefined mixture of six species and their biomasses, including two commonly consumed flatfish species (Solea solea and Pleuronectes platessa) as well as the rare Misgurnus fossilis and three non-native species (i.e., Misgurnus bipartitus, Xenopus laevis, and Lithobates catesbeianus) that are unlikely to occur naturally in the river or to be present in human diets. |
| In the text | |
 |
Fig. 3 Overall treatment barrier effect: Mean eDNA detection probability (±95% CI) of the six spiked species across WWTPs, based on the proportion of positive ddPCR detections before (pre-spike) and after (post-spike) the controlled introduction of the homogenized tissue mixture at the influent. Bars show mean detection probabilities aggregated across all taxa and facilities. The X-axis represents the sampling locations per WWTP, i.e., the influent (incoming untreated wastewater), the effluent (treated discharge water), and the downstream river site (100 m from the effluent outflow). |
| In the text | |
 |
Fig. 4 Measured eDNA concentrations (copies per μL) of the six spiked species (i.e., Xenopus laevis, Misgurnus fossilis, Misgurnus bipartitus, Pleuronectes platessa, Lithobates catesbeianus, and Solea solea) across four WWTPs (Ghent, Ostend, Nevele, and Zomergem) at three sampling locations per WWTP: influent (incoming wastewater), effluent (treated water released from the WWTP), and downstream river (100 m from effluent discharge point). The shaded areas with blender icons indicate the introduction of the DNA spike containing six species. Box plots show the distribution of eDNA concentrations for each species across filter replicates. Note that the Y-axis range differs for each panel. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


