| Issue |
Knowl. Manag. Aquat. Ecosyst.
Number 427, 2026
Conservation genetics
|
|
|---|---|---|
| Article Number | 13 | |
| Number of page(s) | 14 | |
| DOI | https://doi.org/10.1051/kmae/2026006 | |
| Published online | 03 April 2026 | |
Research Paper
Newly discovered diversity within European branchiobdellids – a story of co-evolution and a need for conservation
1
Department of Biology, Faculty of Science, University of Zagreb, Rooseveltov trg 6, Zagreb, Croatia
2
University Hospital for Infectious Diseases “Dr. Fran Mihaljević”, Department for Dangerous Pathogens, Mirogojska 8, Zagreb, Croatia
3
Croatian Natural History Museum, Demetrova 1, Zagreb, Croatia
4
Institute of Biodiversity and Ecosystem Research, Bulgarian Academy of Sciences, 2 Gagarin Street, 1113 Sofia, Bulgaria
* Corresponding author: This email address is being protected from spambots. You need JavaScript enabled to view it.
Received:
12
January
2026
Accepted:
14
March
2026
Abstract
Branchiobdellids are small ectosymbiotic annelids living primarily on freshwater crayfish. Because humans frequently translocate crayfish, these symbionts can be co-translocated, with implications for biosecurity and conservation. Despite over a century of study, relationships between European branchiobdellids and their crayfish hosts (basibionts) remain poorly resolved. Therefore, we analysed 215 mitochondrial COI sequences (67 new and 148 from GenBank and BOLD databases) spanning most documented ectosymbiont–basibiont associations. Phylogenetic reconstruction and species delimitation identified seven recognized species and revealed deep intraspecific divergence in five, suggesting the presence of cryptic lineages. Branchiobdella hexadonta and Branchiobdella parasita each contained multiple genetically distinct groups linked to different crayfish species, whereas Branchiobdella astaci and Branchiobdella balcanica showed strong host-specific monophyly. Branchiobdella italica and Branchiobdella pentadonta showed lower divergence, consistent with a younger origin inferred by molecular clock analyses. Divergence times broadly matched crayfish basibiont history, indicating likely co-speciation. Results support an origin of Branchiobdella inhabiting crayfish gill chambers and show that crayfish translocations may blur natural epibiont–basibiont boundaries, thus complicating monitoring of symbiont lineages. This work advances our understanding of co-evolution in freshwater ecosystems and underscores the conservation importance of preserving both native crayfish and their associated symbionts, whose extinction risks may be underestimated due to unresolved taxonomy.
Key words: Branchiobdella / freshwater crayfish / phylogeny / Europe / mtCOI
Göran Klobučar and Matej Vucić contributed equally to this work.
Deceased.
© G. Klobučar et al., Published by EDP Sciences 2026
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License CC-BY-ND (https://creativecommons.org/licenses/by-nd/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. If you remix, transform, or build upon the material, you may not distribute the modified material.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License CC-BY-ND (https://creativecommons.org/licenses/by-nd/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. If you remix, transform, or build upon the material, you may not distribute the modified material.
1 Introduction
The genus Branchiobdella, belonging to the family Branchiobdellidae, subfamily Branchiobdellinae, represents a group of ectosymbiotic annelids obligatorily associated with freshwater crayfish belonging to the family Astacidae (Holt, 1986; Gelder and Williams, 2016). Native European crayfish are key benthic ecosystem engineers whose numbers are notably decreasing, so they are considered threatened taxa (Jussila et al., 2021). Consequently, though less obviously, the numbers of branchiobdellids, their ectosymbionts, are also irreversibly declining (Szenejko et al., 2023). Branchiobdellids inhabit various microhabitats on their crayfish host (the basibiont), including the carapace and gill chambers, where their interactions with crayfish range from commensalism to mutualism and, sometimes, parasitism (Gelder and Williams, 2016). Because freshwater crayfish are often translocated with their symbionts, understanding branchiobdellid diversity and host association has practical relevance for monitoring, management and conservation of freshwater ecosystems’ biodiversity. The European species of genus Branchiobdella exhibit significant diversity and evolutionary complexity, making their phylogeny a topic of considerable interest in taxonomy, biogeography, and evolutionary biology (Oberkofler et al., 2002; Subchev, 2014; Šarić et al., 2018; Szenejko et al., 2023).
Branchiobdellidans have a Holarctic distribution, with European species forming a distinct clade separate from their North American and East Asian relatives. Modern classifications recognize nine valid species of Branchiobdella in Europe: Branchiobdella astaci Odier, 1823, B. parasita (Braun, 1805), B. pentadonta Whitman, 1882, B. hexadonta Grüber, 1883, B. italica Canegallo, 1928, B. balcanica Moszynsky, 1938, B. kozarovi Subchev, 1978, B. papillosa, and recently described B. bulgariensis Subchev and Rimcheska, 2021. The phylogenetic relationships within Branchiobdella have traditionally relied on morphological traits, such as body size and shape, jaw shape and denticulation, and spermathecal structures (Gelder and Williams, 2016). However, these characters are sometimes insufficient for resolving the species boundaries due to high intraspecific variability and cryptic diversity. Therefore, it has become necessary to employ molecular methods to better understand branchiobdellidan diversity (Gelder and Williams, 2016).
Within Europe, their distribution closely aligns with that of their basibionts, the native freshwater crayfish species (Fig. 1). Their ecological dynamics and evolutionary histories are deeply intertwined with those of their crayfish basibionts (Gelder and Williams, 2016; Šarić et al., 2018; Szenejko et al., 2023). The co-evolution of branchiobdellids and freshwater crayfish is complex, shaped by ecological and evolutionary pressures. When the effects are predominantly beneficial for both the epibiont and basibiont, co-evolution can be expected to lead to associational specificity (Fernandez-Leborans, 2010). Filipiak et al. (2016) emphasize that host-parasite associations are also often characterized by co-speciation, where the phylogenies of the symbionts and hosts show congruent patterns due to long-term interactions. The evolution of symbiotic traits often reflects the environmental conditions and basibiont specificity, as is the case of branchdiobdellids; while many species engage in mutualistic cleaning interactions, others, particularly gill-dwelling species like B. astaci and B. hexadonta, have been associated with parasitic behaviours that can harm their basibionts (Szenejko et al., 2023).
Branchiobdellid species are known to exhibit preferences for specific crayfish species that might be considered their primary basibionts, such as B. balcanica associating with Astacus astacus (Klobučar et al., 2006), B. kozarovi with Pontastacus leptodactylus and A. astacus (Subchev, 2014), or B. italica with Austropotamobius pallipes s.l.. Others, like B. parasita, one of the most widely distributed branchiobdellid species, is most frequently found on A. astacus and Austropotamobius torrentium (Klobučar et al., 2006; Subchev, 2014). Accordingly, the geographic distribution of Branchiobdella also reflects historical biogeographic events that have influenced the evolution of their crayfish basibionts, presumably through allopatric speciation, leading branchiobdellid species to diversify alongside their primary hosts. Preferences for specific crayfish species can (and should) be evaluated by analysing both the abundance and the frequency of occurrence of branchiobdellids on each crayfish species (Klobučar et al., 2006).
Our former studies highlighted the richness of branchiobdellid fauna in Croatia and surrounding countries, underscoring the importance of the Dinarides, a mountain range in south-eastern Europe, and its karstic environment in shaping the evolutionary trajectories of both basibionts and their symbionts (Klobučar et al., 2006; Šarić et al., 2018). However, it is important to highlight that freshwater crayfish, particularly the noble crayfish (A. astacus), have often been translocated by humans because of their high economic value. These human-mediated movements likely also resulted in the translocation of associated branchiobdellid species. If other crayfish species were already present in the new environment, this may have facilitated the transfer of branchiobdellid species to these new local hosts (Bláha et al., 2018). Accordingly, reconstructing branchiobdellid phylogeny can help distinguish long-term associations from recent transfers driven by crayfish movements, supporting interpretation of present-day assemblages.
Recent advances in molecular phylogenetics, particularly the use of mitochondrial cytochrome c oxidase subunit I (COI) sequences, have significantly refined our understanding of Branchiobdella evolution. Molecular evidence from Šarić et al. (2018) revealed high genetic diversity of branchiobdellid species in Croatia and surrounding countries, identifying five main species: B. astaci, B. hexadonta, B. italica, B. parasita, and B. pentadonta and suggests that basibiont-symbiont co-evolution played a crucial role in the diversification of the genus. The congruent phylogenetic patterns observed between European branchiobdellids and their crayfish basibionts in the genus Austropotamobius indicate a history of co-speciation and ecological adaptation, mirroring broader trends in basibiont-symbiont evolution. Additionally, Šarić et al. (2018) demonstrated that molecular analyses could distinguish morphologically cryptic taxa within established species, which displayed divergence in the same manner as their crayfish basibionts. Therefore, additional samples were necessary to gain more resolution of the biogeographic and evolutionary dynamics of the association between the three autochthonous European crayfish species and their branchiobdellid ectosymbionts. A robust phylogenetic framework can support freshwater biodiversity management by detecting introductions and host-switching after crayfish translocations, informing conservation units for native symbiont diversity, and improving screening of crayfish movements.
The aim of the study was to clarify phylogenetic relationships among European branchiobdellid species by analysing the largest dataset assembled to date: mitochondrial COI sequences from specimens collected from European crayfish belonging to the genera Astacus and Austropotamobius, the standard marker used in branchiobdellidan genetic studies. We also aimed to test for cryptic diversification among European branchiobdellid species and to evaluate whether the branchiobdellid evolutionary patterns could provide insights into the evolution of their crayfish basibionts, which in turn could enhance conservation actions dedicated to preserving freshwater biodiversity.
 |
Fig. 1 Locations with sequenced branchiobdellidans (B. astaci, B. balcanica, B. hexadonta, B. italica, B. kozarovi, B. parasita, and B. pentadonta). Areas of distribution for freshwater crayfish: Au. pallipes s.l., Au. torrentium, and A. astacus. A) Europe and B) South-eastern Europe. Crayfish distributions were taken from the IUCN Red List and adapted for this study: A. astacus (Edsman et al., 2010), Au. torrentium (Pârvulescu et al., 2025), and Au. pallipes (Füreder et al., 2010). Maps were produced using QGIS software v. 3.40 (QGIS, 2025). |
2 Materials and methods
Sequences were acquired from Branchiobdella species collected from Astacus astacus, Austropotamobius torrentium and Austropotamobius pallipes from 81 locations, of which 32 examined sites were analysed in this study (Tab. S1). A total of 215 sequences were analysed, comprising 67 obtained in this study and 148 downloaded from the GenBank and The Barcode of Life (BOLD) databases.
2.1 DNA extraction, gene amplification and sequencing
Total genomic DNA was extracted using Macherey-Nagel™ NucleoSpin™ Tissue kit (MN, Germany) following the manufacturer's protocol. PCR was performed to obtain amplicons of the mtDNA marker COI using LCO-1490/HCO-2198 primers (Folmer et al., 1994). Each PCR reaction was carried out in a 20 μL reaction volume containing 0.5-unit HotStarTaq Plus DNA Polymerase (Qiagen, Germany), 200 μM of each dNTP, 1.5 mM MgCl2, 0.25 μM of each primer, and 1 μL of template DNA. PCR cycling conditions followed protocol described in Vitecek et al. (2015).
All obtained amplicons were purified using the ExoAnP enzymatic clean-up (New England Biolabs, USA) following the manufacturer protocol. Sanger sequencing was performed using the Macrogen Europe sequencing service (Amsterdam, Netherlands) using same set of primers as in PCR reactions. Sequencing reactions were conducted using BigDye™ terminator cycling conditions on an ABI 3730XL Automatic Sequencer (Applied Biosystems).
2.2 Data sequencing, phylogenetic analyses and analyses of genetic diversity
Branchiobdella sequences obtained in this study were manually inspected for base-pair ambiguities, indels, and stop codons using BIOEDIT v.7.2.5 (Gene Codes Corporation, Ann Arbor, MI, USA) (Hall, 1999). Subsequently, COI gene sequences were assembled from both forward and reverse reads. In addition to our dataset, publicly available Branchiobdella COI sequences (148) were retrieved from the NCBI GenBank (http://www.ncbi.nlm.nih.gov/nucleotide, accessed on 18/07/2024) and BOLD database for further analyses. All sequences were aligned using the MUSCLE algorithm (Edgar, 2004) in MEGA11 (Tamura et al., 2021). Mesquite ver. 3.5 (Maddison and Maddison 2019) was used to quality-check the alignments. COI haplotypes were defined using Fabox v.1.61 (Villesen, 2007) to minimize redundant information. For downstream phylogenetic analysis, we included only such defined unique sequences or haplotypes.
Using an ultrafast bootstrap approximation (Hoang et al., 2018), the maximum-likelihood tree was inferred using IQ-TREE (Nguyen et al., 2015; Trifinopoulos et al., 2016) while accounting for the optimal substitution model identified by ModelFinder (Kalyaanamoorthy et al., 2017). The TPM2 + F + I + G4 model was applied as the optimal nucleotide substitution model. The sequences of Magmatodrilus obscurus (JQ821600), Triannulata magma (JQ821629) and Hirudo medicinalis (HQ333519) were used as outgroups. Tree was visualized using FigTree v1.4.4 (Rambaut, 2014).
Pairwise distances were calculated in MEGA 11 (Tamura et al., 2021) using the Kimura 2-parameter (K2P) model, with the pairwise deletion option.
Species delimitation analysis was conducted using the multi-rate Poisson Tree Process (mPTP) method (Kapli et al., 2017).
2.3 Divergence time
Based on the results of ML and Bayesian analyses, a data set for divergence time estimation was prepared. It comprised a subset of COI sequences selected to ensure that all revealed subclades were represented by the two most divergent sequences (shaded grey in Tab. S1). The sequences of Magmatodrilus obscurus (JQ821600), Triannulata magma (JQ821629) and Hirudo medicinalis (HQ333519) were used as outgroups. A time-calibrated Bayesian tree, based on the aforementioned subset of COI sequences was estimated using BEAST version 2.7.3 (Bouckaert et al., 2019), available at CIPRES Science Gateway (Miller et al., 2010). The best-fit model of molecular evolution, HKY + G, was selected under Bayesian information criterion (BIC) using JModelTest v.2.1.10 (Darriba et al., 2012). The Yule model was used as the tree prior, and initial analyses were performed under both the strict and optimised relaxed clock. Since the lower value of the 95% HPD interval for ucld.stdev parameter approached zero (value range [1.1272E-10, 1.0486]), indicating minimal rate variation among branches and thus supporting a strict clock assumption (Drummond et al., 2007), for all further analyses strict clock model was used. Several different calibrations approaches were applied. Molecular clock rate was estimated using different calibration points alone or combined: 1) the split of Au. torrentium and Au. pallipes s.l. (Dinarides uplift 16 million years ago (MYA) (Jelić et al., 2016) and Dinarides uplift 12 MYA (Klobučar et al., 2013)), 2) the split of North American and European astacids (66.22 MYA (BPBP, Brackem-Grisson et al., 2014)), 3) two-point calibration (66.22 and 16 MYA and 66.22 and 12 MYA). The final molecular clock rate value used for divergences dating was based on the range (minimum and maximum) rate values obtained by calibrations 1-3. Analyses were conducted using Markov Chain Monte Carlo (MCMC) chains run for 10 million generations, sampling every 1000 generations. The convergence of the runs was assessed based on ESS values and trace plots in Tracer v. 1.7.2 (Rambaut et al., 2018), and the first 10% of the sampled trees were discarded as burn-in.
3 Results
3.1 Sampling, DNA extraction, gene amplification and sequencing
After genomic DNA extraction and sequence quality examination, we obtained 67 new, high-quality COI sequences belonging to 6 branchiobdellid species from specimens collected in Bulgaria, Croatia, Lithuania, and Montenegro (Fig. 1, Fig. S1). Of these, 61 were unique haplotypes (Tab. S1). These sequences were compared with 148 European branchiobdellid sequences available in the NCBI GenBank and BOLD, including 106 unique haplotypes. Altogether, 215 sequences belonging to 7 branchiobdellid species were included in the analyses (Fig. 1, Tab. 1, Fig. S1, Tab. S1). The specimens were collected from their respective basibionts Au. pallipes s.l., Au. torrentium, and A. astacus in Austria, Bulgaria, Croatia, Czechia, Germany, Italy, Lithuania, Montenegro, Netherlands, North Macedonia, Poland, and UK. For phylogenetic reconstruction under the maximum likelihood and for divergence time estimation, we employed a dataset comprising 167 defined haplotypes. Sequences for two branchiobdellid species were not obtained, B. papillosa, which is known only from its type locality at Voralberg, Austria (Nesemann and Hutter, 2002) where attempts to collect new material were not successful (Subchev, 2014), and for the recently described B. bulgariensis from Bulgaria (Subchev and Rimcheska, 2021). Sequences for the other seven species were obtained from individuals found on their respective basibionts (Tab. 1), except for B. kozarovi that was not obtained from P. leptodactylus but from A. astacus. In Table 1 we have marked all the crayfish basibiont species for each of the nine European branchiobdellid species according to the existing literature data (Oberkofler et al., 2002; Klobučar et al., 2006; Subchev, 2014; Vlach et al., 2017; Šarić et al., 2018; Shrestka and Utevsky, 2024). In this study, we have managed to obtain sequences (grey shading) for seven branchiobdellid species found on their respective known basibiont species. Epibiont-basibiont associations that are rarely documented in the literature and are presumed to be either misidentified or are a result of a recent transfer between crayfish species were indicated with question marks.
Crayfish basibiont species and European branchiobdellid species. (+ confirmed findings in the literature, ? – rare findings – possible transfer from other crayfish species or misidentification, cells with grey background – species sequenced in this study).
3.2 Phylogenetic analysis, species delimitation, and genetic diversity
Phylogenetic reconstructions based on Maximum Likelihood (ML) and Bayesian Implementation methods revealed strong bootstrap support for major clades within the Branchiobdella (Fig. 2). The seven recognized Branchiobdella species (B. astaci, B. parasita, B. pentadonta, B. hexadonta, B. italica, B. kozarovi, and B. balcanica) formed distinct clusters, with varying levels of support for their monophyly. The sister-group relationship of B. astaci and B. parasita, as well as that of B. pentadonta and B. italica (the latter being part of the B. pentadonta complex) was well supported, suggesting a close evolutionary relationship between these species. Similarly, B. kozarovi and B. balcanica also exhibited a close genetic relationship. Also, the presence of additional genetic structuring within several Branchiobdella species suggests the existence of undescribed lineages that are a consequence of geographic variation or basibiont-specific adaptations, or both.
Phylogenetic analyses revealed distinct COI groups within previously known species and, combined with the results of species delimitation methods, suggest the existence of new and so far undescribed species. In the mPTP analysis for all values of prior maximal distance, initial partitions identified 15 groups (Fig. 2). Three groups were found within B. pentadonta (4, 5, and 6), three within B. hexadonta (9, 10, and 11), and three corresponded to B. parasita (13, 14, and 15). Two groups were found within B. kozarovi (1 and 2), and within B. italica (7 and 8), while B. balcanica and B. astaci both consisted of a single group (3 and 12, respectively).
The monophyly of B. astaci (group 12, on Au. pallipes s.l.) and B. balcanica (group 3, on A. astacus) was well supported, while the monophyly of the remaining species was only weakly to moderately supported by bootstrap values. The monophyly of B. hexadonta was not confirmed by ML analysis. Sequences assigned to this species clustered into three well-supported subclades/groups belonging to the three different basibiont species: Au torrentium (group 11), Au. pallipes s.l. (group 9 represented by one sequence), and A. astacus (group 10). Similarly, B. pentadonta was structured into three well supported groups where first (4) was attributed in large part to A. astacus, and the other two (5, 6) to Au. torrentium. However, in subclade 5 one of the two branchiobdellid sequences came from an unknown basibiont. Furthermore, B. parasita, represented by the highest number of sequences, was also divided into three groups (13, 14, and 15) that varied in their level of statistical support: group nr. 14 represented by a single sequence (A. astacus basibiont, pers. comm.), group 13 presented by B. parasita specimens collected from Au. pallipes s.l., and the largest subclade, nr. 15 comprised of B. parasita collected from three different basibionts (Au. torrentium, Au. pallipes s.l., and A. astacus). In contrast, B. kozarovi showed strongly supported genetic structuring into two groups (1 and 2), on A. astacus basibiont. Similarly, B. italica exhibited two well-supported groups (7 and 8), coming mainly from one basibiont species – Au. pallipes s.l. Branchiobdella italica has also been occasionally found on Au. torrentium (Tab. 2, Fig. 2) that are within or close to the distribution area of Au. pallipes s.l. and are believed to be the result of past translocations.
Interspecific and intraspecific divergence between sequence pairs for groups recognized by mPTP analysis was estimated and is presented in Tables. 2–8. Obtained interspecific p-distance values among the seven analysed branchiobdellid species ranged from 12.7% between sister groups B. italica and B. pentadonta to 25.6% between B. pentadonta and B. kozarovi (Tab. 2). For the groups defined by species delimitation, p-distances ranged from 0.08% to 8.03% (Tab. 3), and their geographical distribution is shown in Figure 2.
For B. hexadonta, p-distances ranged from 17.3% between groups 9 and 11 to 23.9% between groups 9 and 10. For B. parasita, the values range from 12.9% between groups 13 and 15 to 18.7% between groups 13 and 14. In the case of B. kozarovi, the obtained value between groups 1 and 2 was 12.4%. Somewhat lower values were observed for groups in B. pentadonta (4.1 to 9%) and B. italica (8.6%).
 |
Fig. 2 The maximum-likelihood tree of genus Branchiobdella based on cytochrome c oxidase subunit I gene analysis was inferred with IQ-Tree (Nguyen et al., 2015). The numbers on the branches present ultrafast bootstrap values (≥0.7). Sequences were annotated with haplotype number according to Table S1. As presented in the legend, the symbols next to the haplotype names indicate the crayfish host. Asterisks (*) represent the new haplotypes obtained in this study. Groups recognized by mPTP analysis are displayed on the right side of the phylogram before the branchdiobdellidan species affiliation or name. |
Inter- and intraspecific divergence over sequence pairs for Branchiobdella species. Average and range values (minimum and maximum values are shown in parentheses) of uncorrected p-distances among and within studied Branchiobdella species are presented.
Intraspecific divergence over sequence pairs for groups recognized by mPTP analysis. P-distances for groups defined by using species delimitation method (NA – not applicable, only one sequence).
3.3 Divergence time
The estimated divergence times for the 15 groups revealed by species delineation analysis are shown in Figure 3. As expected, the initial split accounted for the divergence between North American and European Branchiobdellida species is approximately 48 million years ago (MYA). Within European species, the first moderately supported split refers to separation of the group comprising B. hexadonta, B. astaci and B. parasita from the group containing B. kozarovi, B. balcanica, B. pentadonta and B. italica that occurred around 41 MYA. Subsequent splitting within the former group occurred during the 23-29 MYA interval, while divergence within the latter group was estimated at 15-23 MYA. However, none of above groups received support from Bayesian posterior probabilities. Rather, the results suggested an unresolved tetratomy among B. hexadonta, B. astaci + B. parasita, B. kozarovi + B. balcanica and B. pentadonta + B.italica. The very similar divergence times and overlapping 95% HPD intervals suggested parallel evolution of B. hexadonta and B. astaci + B. parasita. A similar pattern was observed for the most recent species pair, B. pentadonta and B. italica, whose estimated divergence time overlap, at approximately 5-7 MYA.
 |
Fig. 3 Time-calibrated Bayesian phylogenetic tree inferred from mitochondrial COI sequences. The divergence times are depicted as median values and 95% HPD intervals. Circles on nodes represent clade support (Bayesian posterior probabilities, PP) as shown in the figure. 15 groups revealed by species delimitation are shown as well as employed haplotypes (details are provided in Tab. S1). Further, the names of branchiobdellid species are provided next to branches. Nodes A and B represent calibration points: A – the split of North American and European Astacids, and B – the split of Au. torrentium and Au. pallipes s.l. |
4 Discussion
4.1 Phylogenetic analysis and genetic diversity
This study added 67 new sequences (61 unique haplotypes) of the European Branchiobdella species, increasing the number of available sequences by 37% and revealing greater diversity than previously known. Seven recognized Branchiobdella species (B. astaci, B. parasita, B. pentadonta, B. hexadonta, B. italica, B. kozarovi, and B. balcanica) formed distinct clusters in a phylogenetic analysis (Fig. 2). The close genetic relationship between B. kozarovi and B. balcanica, as evidenced by their branching from other branchiobdellid species, aligns with their shared basibiont, A. astacus. This is not surprising since the two branchiobdellid species are considered primary ectosymbionts of closely related genera Astacus and Pontastacus, which show a similar divergence pattern from their common ancestor with Austropotamobius (Maguire et al., 2014; Crandall and De Grave, 2017; Lovrenčić et al., 2020; Bláha et al., 2023). These results indicate that A. astacus and P. leptodactylus are indeed primary basibionts for B. balcanica and B. kozarovi, respectively (Subchev, 2014). This finding supports co-evolution of these branchiobdellid species with their original crayfish basibionts A. astacus and P. leptodactylus (formerly Astacus leptodactylus) whose distributions are centred mainly in Central and Eastern Europe, respectively (Kouba et al., 2014).
The sister-group relationship of B. italica and B. pentadonta (part of the B. pentadonta complex, Karaman, 1970; Šarić et al., 2018) is also evident in this phylogenetic analysis, where they formed distinct and a well-supported cluster that suggests a close evolutionary relationship between these two species. Branchiobdella italica is almost exclusively present on Au. pallipes s.l. whereas the morphologically very similar sister species B. pentadonta is associated with Au. torrentium and A. astacus. Knowing that Au. pallipes s.l. and Au. torrentium are, according to phylogenetic analyses and divergence time estimates, relatively young sister species (Trontelj et al., 2005; Klobučar et al., 2013; Jelić et al., 2016; Lovrenčić et al., 2020) it can be assumed that sister species B. italica and B. pentadonta evolved on these crayfish species as suggested by Šarić et al. (2018). Lower values of evolutionary divergence (p-distances) between these two branchiobdellid species as well as of their primary basibionts support this conclusion (Trontelj et al., 2005; Šarić et al., 2018).
It is also worth noting that the jaws of B. pentadonta are similar in shape and dentation to those of B. italica (Subchev, 2014), as well as the primary locations of their cocoons on crayfish hosts (Shrestkha et al., 2025), further supporting their close phylogenetic relationship. Subchev (2014) also highlights the similarity of B. pentadonta's jaws to those of B. balcanica, and in terms of dentation, to those of B. kozarovi. Branchiobdella pentadonta has been more frequently found on Au. torrentium then on A. astacus (Klobučar et al., 2006; Shrestkha et al., 2025), further supporting the stone crayfish is its primary basibiont. The occurrence of B. pentadonta on A. astacus likely reflects overlap between the ranges of Au. torrentium and A. astacus in Central and Eastern Europe, leading to secondary contact after divergence, and/or the introduction of A. astacus into Au. torrentium habitats (Maguire et al., 2013).
The third cluster has an unresolved dichotomy and indicates close relationship of B. hexadonta, and B. astaci + B. parasita. Results of our study showed that B. astaci is almost exclusively found on Au. pallipes s.l., while B. hexadonta and B. parasita were collected from all three crayfish species. Branchiobdella astaci has also been reported on A. astacus in Eastern Europe and on Au. torrentium in Bulgaria (Subchev, 2014). These three branchiobdellid species, along with B. pentadonta, have similar distribution in Europe (Subchev, 2014; Shrestkha and Utevsky, 2024; Subchev et al., 2024) and are therefore difficult to associate with their primary basibionts.
To elucidate co-evolution between branchiobdellid species and their crayfish basibionts, it is important to identify the original epibiont-basibiont associations, as allopatric co-speciation likely occurred (Filipiak et al., 2016). This is challenging because anthropogenic introductions and translocations, together with natural range expansion of crayfish species, can obscure original host associations. Nevertheless, several studies have reported the prevalence of particular branchiobdellid species on specific crayfish hosts and documented these patterns of association (Klobučar et al., 2006; Shrestkha and Utevsky, 2024; Konno, 2025; Shrestkha et al., 2025). These data are certainly useful in determining the preference of branchiobdellids toward their primary basibiont species. For example, it has been confirmed that B. kozarovi is found mainly on P. leptodactylus and B. balcanica on A. astacus (Subchev, 2014; Shrestkha and Utevsky, 2024). Certain branchiobdellid species have been discovered on several crayfish species but it is evident that sometimes they appear in lower numbers what that be the result of a transfer from the primary basibiont species, e.g. lower abundance of B. pentadonta on A. astacus in comparison to B. balcanica or B. parasita (Klobučar et al., 2006; Shrestkha and Utevsky, 2024).
4.2 Species delimitation and divergence time
Divergence time estimates indicated that split between North American and European branchiobdellidans occurred 36.8–59.9 MYA, which, expectedly, coincides with the separation of North American and European crayfish from the family Astacidae (Bracken-Grissom et al., 2014), and is similar to observations made by Konno (2025) in the study of Japanese branchiobdellidans. For European branchiobdellids molecular clock analyses (Fig. 3) suggest that three closely related species: B. hexadonta, B. astaci, and B. parasita, represent the earliest lineages within the European group. The gill-dwelling B. hexadonta appears to be the oldest, diverging from the common ancestor of B. astaci (also gill-dwelling) and B. parasita approximately 28–44 million years ago, with the latter two splitting around 22–37 million years ago. Based on these findings, it can be hypothesized that the earliest branchiobdellid species (or their ancestors) likely inhabited the gill chambers of crayfish, a habitat that is both easily accessible and offers protection as well as abundant food resources.
Branchiobdella hexadonta exhibited the highest observed intraspecific p-distance values, attributed to the presence of three well-supported groups (9, 10, and 11), each showing high levels of evolutionary divergence (mean 17–24%, Tab. 4) comparable to or exceeding interspecific distances among recognized branchiobdellid species (Tab. 2). These elevated intraspecific divergences suggest the presence of cryptic taxa (“hexadonta-complex”) and highlight the need for detailed morphological reassessment of the specimens from these groups. Each group was associated with different crayfish species (Au. torrentium – group 11, Au. pallipes s.l. – group 9, and A. astacus – group 10) and corresponds to the distribution areas of the matching basibionts (Fig. 4).
Šarić et al. (2018) also observed three groups within B. hexadonta (two on Au. torrentium and one on A. astacus) but in the present research B. hexadonta from Au. torrentium forms a single group (11) and a new group (9) from a novel basibiont (Au. pallipes s.l.) was detected. The observed genetic divergence was greater between B. hexadonta groups associated with different crayfish genera (i.g., Astacus and Austropotamobius) than among groups occurring on hosts within the same genus (Au. torrentium and Au. pallipes s.l.) supporting co-evolution between the epibionts and their basibionts.
The pronounced separation among groups and their respective basibionts may reflect the gill-dwelling lifestyle of B. hexadonta, which might restrict (but not prevent) host-switching. Given the observed deep divergences, especially for group 10 with basibiont A. astacus (21-35 MYA), it is plausible that the common ancestor of B. hexadonta complex was already present in the gill chambers of the common crayfish ancestor of Astacus/Pontastacus and Austropotamobius.
Another gill-dwelling branchiobdellid – B. astaci, which has never been reported co-occurring with B. hexadonta on the same basibiont (Subchev, 2014) was found almost exclusively in the gill chambers of Au. pallipes s.l. (group 12) from two geographically distant locations: Istria (Croatia) and Yorkshire (UK). These findings lend support to the hypothesis that Au. pallipes s.l. was anthropogenically translocated from the Mediterranean region to the United Kingdom via France during the Middle Ages (Kouba et al., 2014). This scenario is also supported by the observed high haplotype diversity of B. astaci in Croatia (Tab. S1) compared to the UK. There is only one B. astaci haplotype (Hap 117) that was collected from A. astacus in Lithuania and is very similar to haplotypes from Yorkshire, UK, possibly indicating transfer of B. astaci from Au. pallipes s.l. to A. astacus. In Klobučar et al. (2006), the only finding of B. astaci in A. astacus gills was in Istria, Croatia, where A. astacus has been introduced and Au. pallipes s.l. occurs naturally (Maguire et al., 2011) with B. astaci frequently present in its gill chambers. Findings of B. astaci on Au. torrentium and P. leptodactylus are scarce (Subchev et al., 2024) and thus can be considered as a consequence of transfer from their primary basibiont-crayfish species or a misidentification.
Branchiobdella parasita is closely related to B. astaci (Figs. 2 and 3), a relationship also reflected in the similarity of their triangular jaw morphology. It is therefore tempting to hypothesize that B. parasita evolved from the gill-dwelling ancestor of these two species that consequently conquered the exoskeleton of the crayfish. This is probably indicating ecological speciation or its combination with allopatric speciation. Using the species delimitation method, B. parasita, the most common and widespread European branchiobdellid, was resolved into three distinct groups according to mPTP analysis (13, 14 and 15, Fig. 5) as already reported by Šarić et al. (2018).
Szenejko et al. (2023) also found a high genetic distance p-value between B. parasita specimens obtained from A. astacus in Germany and Poland and those collected from Au. torrentium (5.9%) and Au. pallipes s.l. (10.2%) and have suggested the existence of “parasita-complex” that demands its taxonomic revision. In our study, we analyzed a larger number of sequences, and the observed p-distances of 13–19% exceed the divergence values reported between the youngest species, B. italica and B. pentadonta (12.7%) (Tabs. 2 and 5), supporting the presence of cryptic taxa. Branchiobdella italica and B. pentadonta have also been previously considered as part of “Branchiobdella pentadonta-complex” (Šarić et al., 2018). Two smaller groups within B. parasita (13 and 14) were each associated with a single basibiont species: Au. palipes s.l. and A. astacus, respectively, and, as in B. hexadonta groups, genetic divergence value was higher for groups having basibionts from different crayfish genera (Astacus and Austropotamobius). Larger sample sizes for these groups would be more informative as whether these groups are linked to a single basibiont species or, as in the case of the largest group (15), that included large number of new haplotypes, to different basibiont species (Au. torrentium, Au. pallipes s.l., and A. astacus). However, it is important to emphasize that the Dinaric karst region of Croatia and Slovenia seems to constitute the hotspot of genetic diversity for B. parasita (Fig. 5). Notably, this region also harbours the highest genetic diversity in Au. torrentium and Au. pallipes (Klobučar et al., 2013; Jelić et al., 2016; Lovrenčić et al., 2020) confirming its importance as the most probable region of speciation of the Austropotamobius and its centre of radiation. The broader host association of B. parasita complicates the determination of its primary basibiont, likely reflecting interspecific crayfish contacts and anthropogenic translocations, especially of the economically important A. astacus. B. parasita inhabits the surface of the crayfish exoskeleton and can therefore be relatively easily transferred to another crayfish during the physical contact (Bláha et al., 2018; Parpet and Gelder, 2020). Moreover, it should be emphasized that A. astacus has been the most frequent basibiont for B. parasita.
The observed p-distances among the remaining groups – three in B. pentadonta (groups 4, 5, and 6; Fig. S1), two in B. italica (groups 7 and 8, Fig. S2), and two in B. kozarovi (groups 1 and 2) – along with the estimated divergence times (Fig. 3), are lower than the divergence between recognized species B. italica and B. pentadonta (12.7%) (Tabs. 2, 6, 7, and 8). Notably, groups of B. italica and B. kozarovi were associated with one host species, Au. pallipes s.l. and A. astacus, respectively. If Au. pallipes s.l. is, based on the available data, considered the primary basibiont of B. italica and Au. torrentium of B. pentadonta, the relatively low genetic divergence between their ectosymbiont groups becomes understandable. This pattern reflects the comparatively recent origin of Austropotamobius species (approx. 8-18 MYA; Lovrenčić et al., 2020), which is consistent with the estimated divergence time of B. italica and B. pentadonta in this study (approx. 11-19 MYA; Lovrenčić et al., 2020). In B. pentadonta, groups 5 and 6 are associated with Au. torrentium and are more closely related to each other than to group 4, which occurs almost exclusively on A. astacus – only two records are from Au. torrentium, and one from Au. pallipes s.l.. In the study of Šarić et al. (2018) where branchiobdellids were obtained only from crayfish belonging to genus Austropotamobius no groups of B. pentadonta were determined with the species delimitation method. However, in the same study, the groupings within B. pentadonta mirrored three phylogroups observed in Au. torrentium populations: Gorski Kotar “GK”, Žumberak, Plitvice, Bjelolasica “ŽPB” (from the northern-central Dinaric region), and Southern Balkans “SB” (Klobučar et al., 2013, Lovrenčić et al., 2020). In the current study, while group 5 is confirming the existence of the “SB” phylogroup, group 6 includes B. pentadonta specimens found on Au. torrentium belonging to “GK”, “ŽPB” and Lika and Dalmatia “LD” phylogroups as defined in Klobučar et al. (2013) and Lovrenčić et al. (2020). As one of the youngest species in the genus Branchiobdella, the observed groups within B. pentadonta may not yet have diversified into distinct species; rather, they currently represent lineages co-evolving with their basibionts, Au. torrentium in the Dinaric karst and southern Balkans, and A. astacus. The association of B. pentadonta group 4 with A. astacus most likely arose through host transfer or switching during the first contact between the two crayfish species, approximately 5 MYA (Fig. 3), when Au. torrentium expanded eastward and A. astacus westward (Klobučar et al., 2013; Lovrenčić et al., 2020; Bláha et al., 2023). In order to provide a better explanation of divergence within B. kozarovi, additional sampling, especially from P. leptodactylus, is needed.
 |
Fig. 4 Distribution of three well-supported groups (9, 10, and 11) within B. hexadonta. |
Interspecific divergence over sequence pairs for groups within B. hexadonta recognized by mPTP analysis. Basibionts of specific groups are indicated.
Interspecific divergence over sequence pairs for groups within B. parasita recognized by mPTP analysis. Basibionts of specific groups are indicated.
Interspecific divergence over sequence pairs for groups within B. pentadonta recognized by mPTP analysis. Basibionts of specific groups are indicated.
Interspecific divergence over sequence pairs for groups within B. italica recognized by mPTP analysis. Basibionts of specific groups are indicated.
Interspecific divergence over sequence pairs for groups within B. kozarovi recognized by mPTP analysis. Basibionts of specific groups are indicated.
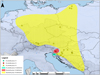 |
Fig. 5 Distribution of three well-supported groups (13, 14, and 15) within B. parasita. |
4.3 Conclusion
Phylogenetic analyses confirmed the existence of seven recognized Branchiobdella species with deep intraspecific divergence within five of them. We confirmed the majority of groups from Šarić et al. (2018) detected within B. parasita, B. hexadonta, and B. italica, and found new groups within B. hexadonta, B. pentadonta, and B. kozarovi. Groups within B. hexadonta and B. parasita show intraspecific genetic divergences that are equal or higher than those between recognized branchiobdellan species and therefore indicate the presence of cryptic taxa and species complexes that necessitate future taxonomic revision. Branchiobdellid species (B. astaci, B. balcanica and B. kozarovi) and groups within species (B. hexadonta, B. italica) are associated with specific crayfish basibionts implying their co-evolution. However, anthropogenic factors (species misidentification, reallocation and introduction of crayfish species) as well as natural contact zones between crayfish make it difficult to trace co-evolution of branchiobdellids with their primary crayfish basibionts. The largest of the three B. parasita groups, showed less host specificity, appearing on all three studied crayfish species (Au. pallipes s.l., Au. torrentium, and A. astacus). Interestingly, two of the three earliest-diverging species, B. hexadonta and B. astaci, are gill-dwellers, which may indicate that the genus Branchiobdella originated within crayfish gill chambers. These new data provide a reference for monitoring and conserving crayfish species and their branchiobdellid ectosymbionts. The decline of European freshwater crayfish due to habitat degradation, invasive species and the crayfish plague (Maguire et al., 2018) also threatens their obligate ectosymbionts, autochthonous European branchiobdellids (Parpet and Gelder, 2020). Loss of host/basibiont populations risks erasing co-evolutionary history and genetic diversity of Branchiobdella spp.
Acknowledgments
This research did not receive any specific grant. All the materials have been collected under approved ethics guidelines and no collection permits were needed.
Supplementary Material
Figure S1. Distribution of three groups (4, 5, and 6) within B. pentadonta.
Figure S2. Distribution of two groups (7 and 8) within B. italica.
Table S1. Species delimitation group.
Access Supplementary MaterialReferences
- Bláha M, Ložek F, Buřič M, Kouba A, Kozák P. 2018. Native European branchiobdellids on non-native crayfishes: Report from the Czech Republic. J Limnol 77: 164–168. [Google Scholar]
- Bláha M, Patoka J, Policar T, Śliwińska K, Alekhnovich A, Berezina N, Petrescu AM, Mumladze L, Weiperth A, Jelic M, Kozák P, Maguire I. 2023. Phylogeographic patterns of genetic diversity in Pontastacus leptodactylus (Decapoda: Astacidae): is the hypothesis of the taxonomically rich genus Pontastacus true? J Linn Soc London Zool 199: 140–155. [Google Scholar]
- Bouckaert R, Vaughan TG, Barido-Sottani J, Duchêne S, Fourment M, Gavryushkina A, Heled J, Jones G, Kühnert D, De Maio N, Matschiner M, Mendes F.K, Müller NF, Ogilvie HA, du Plessis L, Popinga A, Rambaut A, Rasmussen D, Siveroni I, Suchard MA, Wu C-H, Xie D, Zhang C, Stadler T, Drummond AJ. 2019. BEAST 2.5: An advanced software platform for Bayesian evolutionary analysis. PLoS Comput Biol 15: e1006650. [Google Scholar]
- Bracken-Grissom HD, Ahyong ST, Wilkinson RD, Feldmann RM, Schweitzer CE, Breinholt JW, Bendall M, Palero F, Chan TY, Felder DL, Robles R, Chu KH, Tsang LM, Kim D, Martin JW, Crandall KA. 2014. The emergence of lobsters: phylogenetic relationships, morphological evolution and divergence time comparisons of an ancient group (Decapoda: Achelata, Astacidea, Glypheidea, Polychelida). Syst Biol 63: 457–479. [Google Scholar]
- Crandall KA, De Grave S. 2017. An updated classification of the freshwater crayfishes (Decapoda: Astacidea) of the world, with a complete species list. J Crustac Biol 37: 615–653. [Google Scholar]
- Darriba D, Taboada GL, Doallo R, Posada D. 2012. jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9: 772–772. [CrossRef] [PubMed] [Google Scholar]
- Drummond AJ, Ho SYW, Rawlence N, Rambaut A. 2007. A rough guide to BEAST 1.4. University of Edinburgh, Edinburgh, 1–41. [Google Scholar]
- Edgar RC. 2004. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797. [Google Scholar]
- Edsman L, Füreder L, Gherardi F, Souty-Grosset C. 2010. Astacus astacus. The IUCN Red List of Threatened Species 2010: e.T2191A9338388. https://doi.org/10.2305/IUCN.UK.2010-3.RLTS.T2191A9338388.en. Accessed on 17 September 2025 [Google Scholar]
- Fernandez-Leborans G. 2010. Epibiosis in Crustacea: an overview. Crustaceana 83: 549–640. [CrossRef] [Google Scholar]
- Filipiak A, Zając K, Kűbler D, Kramarz P. 2016. Coevolution of host-parasite associations and methods for studying their cophylogeny. Invertebrate Surviv J 13: 56–65. doi: https://doi.org/10.25431/1824-307X/isj.v13i1.56-65 [Google Scholar]
- Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. 1994. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3: 294–299. [PubMed] [Google Scholar]
- Füreder L, Gherardi F, Holdich D, Reynolds J, Sibley P, Souty-Grosset C. 2010. Austropotamobius pallipes. The IUCN Red List of Threatened Species 2010: e.T2430A9438817. https://doi.org/10.2305/IUCN.UK.2010-3.RLTS.T2430A9438817.en. Accessed on 17 September 2025 [Google Scholar]
- Gelder SR. 1996. A review of the taxonomic nomenclature and a checklist of the species of the Branchiobdellae (Annelida: Clitellata). Proc Biol Soc Wash 109: 653–663. [Google Scholar]
- Gelder SR, Williams BW. 2016. Global overview of Branchiobdellida. In Freshwater Crayfish: A Global Overview, CRC Press, 628–654. [Google Scholar]
- Hall TA. 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41: 95–98. [Google Scholar]
- Hoang DT, Chernomor O, Von Haeseler A, Minh BQ, Vinh LS. 2018. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol Biol Evol 35: 518–522. [Google Scholar]
- Holt PC. 1986. Newly established families of the order Branchiobdellida (Annelida: Clitellata) with a synopsis of the genera. Proc Biol Soc Wash 99: 676–702. [Google Scholar]
- Jelić M, Klobučar GIV, Grandjean F, Puillandre N, Franjević D, Futo M, Amouret J, Maguire I. 2016. Insights into the molecular phylogeny and historical biogeography of the white-clawed crayfish (Decapoda, Astacidae). Mol Phylogenet Evol 103: 26–40. [CrossRef] [PubMed] [Google Scholar]
- Jussila J, Edsman L, Maguire I, Diéguez-Uribeondo J, Theissinger K. 2021 Money Kills Native Ecosystems: European Crayfish as an Example. Front Ecol Evol 9: 648495. [CrossRef] [Google Scholar]
- Kalyaanamoorthy S, Minh BQ, Wong TKF, Von Haeseler A, Jermiin LS. 2017. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14: 587–589. [Google Scholar]
- Kapli P, Lutteropp S, Zhang J, Kobert K, Pavlidis P, Stamatakis A, Flouri T. 2017. Multi-rate Poisson tree processes for single-locus species delimitation under maximum likelihood and Markov chain Monte Carlo. Bioinformatics 33: 1630–1638. [Google Scholar]
- Karaman SM. 1970. Beitrag zur Kenntnis der europaischen Branchiobdelliden (Clitellata, Branchiobdellidea). Int Rev Hydrobiol 55: 325–333. [Google Scholar]
- Klobučar GIV, Maguire I, Gottstein S, Gelder SR. 2006. Occurrence of Branchiobdellida (Annelida: Clitellata) on freshwater crayfish in Croatia. Ann Limnol - Int J Lim 42: 251–260. [Google Scholar]
- Klobučar G, Podnar M, Jelić M, Franjević D, Faller M, Štambuk A, Gottstein S, Simić V, Maguire I. 2013. Role of the Dinaric Karst (western Balkans) in shaping the phylogeographic structure of the threatened crayfish Austropotamobius torrentium. Freshwater Biol 58: 1089–1105. [Google Scholar]
- Konno T. 2025. Past, present, and future of Japanese branchiobdellidans: insights into evolutionary history and conservation. PhD thesis, Hokkaido University Collection of Scholarly and Academic Papers, 142 p. [Google Scholar]
- Kouba A, Petrusek A, Kozák P. 2014. Continental-wide distribution of crayfish species in Europe: update and maps. Knowl Manag Aquat Ecosyst 413: 05. [CrossRef] [EDP Sciences] [Google Scholar]
- Lovrenčić L, Bonassin L, Boštjančić LL, Podnar M, Jelić M, Klobučar G, Jaklič M, Slavevska-Stamenković V, Hinić J, Maguire I. 2020. New insights into the genetic diversity of the stone crayfish: taxonomic and conservation implications. BMC Evol Biol 20: 1–20. [CrossRef] [PubMed] [Google Scholar]
- Maguire I, Jelić M, Klobučar G. 2011. Update on the distribution of freshwater crayfish in Croatia. Knowl Manag Aquat Ecosyst 401: 31. [Google Scholar]
- Maguire I, Podnar M, Jelić M, Štambuk A, Schrimpf A, Schulz H, Klobučar G. 2014. Two distinct evolutionary lineages of the Astacus leptodactylus species-complex (Decapoda: Astacidae) inferred by phylogenetic analyses. Invertebr Syst 28: 117–123. [CrossRef] [Google Scholar]
- Maguire I, Klobučar G, Žganec K, Jelić M, Lucić A, Hudina S. 2018. Recent changes in distribution pattern of freshwater crayfish in Croatia − threats and perspectives. Knowl Manag Aquat Ecosyst 419: 2. [CrossRef] [EDP Sciences] [Google Scholar]
- Maguire I, Špoljarić I, Klobučar G. 2013. The indigenous crayfish of Plitvice Lakes National Park, Croatia. Freshwater Crayfish 19: 91–96. [Google Scholar]
- Maddison WP, Maddison DR. 2019. Mesquite: a modular system for evolutionary analysis. Version 3.5. Available at https://www.mesquiteproject.org [Google Scholar]
- Miller M, Pfeiffer W, Schwartz T. 2010. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. Proceedings of the Gateway Computing Environments Workshop (GCE). IEE, New Orleans, 1–8. [Google Scholar]
- Nesemann H, Hutter G. 2002. Krebsegel (Branchiobdellidae: Branchiobdella ODIER, 1823) in Vorarlberg (Österreich) mit einer Neubeschreibung von Branchiobdella papillosa n. sp. Vorarlberger Naturschau 11: 203–214. [Google Scholar]
- Nguyen L-T, Schmidt HA, Von Haeseler A, Minh BQ. 2015. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol Biol Evol 32: 268–274. [Google Scholar]
- Oberkofler B, Quaglio F, Füreder L, Fioravanti M.L, Giannetto S, Morolli C, Minelli G. 2002. Species of Branchiobdellidae (Annelida) on freshwater crayfish in south Tyrol (northern Italy). Bull Fr Pêche Piscic 367: 777–784. [Google Scholar]
- Parpet JF, Gelder SR. 2020. North American Branchiobdellida (Annelida: Clitellata) or Crayfish Worms in France: the most diverse distribution of these exotic ectosymbionts in Europe. Zoosymposia 17: 121–140. [CrossRef] [Google Scholar]
- Pârvulescu L, Ion M, Ács AR. 2025. Austropotamobius torrentium. The IUCN Red List of Threatened Species 2025: e.T260451059A260451126. https://doi.org/10.2305/IUCN.UK.2025-1.RLTS.T260451059A260451126.en. Accessed on 17 September 2025. [Google Scholar]
- QGIS.org (2025.) QGIS Geographic Information System. QGIS Association. http://www.qgis.org [Google Scholar]
- Rambaut A. 2014. FigTree, Version 1.4.4. Available online: http://tree.bio.ed.ac.uk/software/figtree. Accessed on 07 April 2025. [Google Scholar]
- Rambaut A, Drummond A.J, Xie D, Baele G, Suchard MA. 2018. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 675: 901–904. [Google Scholar]
- Šarić I, Klobučar G, Podnar M, Štambuk A, Maguire I. 2018. Molecular phylogeny of branchiobdellidans (Annelida: Clitellata) and their host–epibiont association with Austropotamobius freshwater crayfish. Invertebr Syst 32: 55–68. [Google Scholar]
- Shrestkha M, Utevsky S. 2024. The distribution of branchiobdellidan worms (Annelida: Clitellata) on the noble crayfish, Astacus astacus, in the Transcarpathian region, Ukraine. J VN Karazin Kharkiv Natl Univ Ser Biol 43: 85–95. [Google Scholar]
- Shrestkha M, Utevsky S, Trontelj P. 2025. Diversity, distribution and host interactions of branchiobdellidans (Clitellata: Branchiobdellida) in the Western Balkans. Nat Slov 27: 21626. [Google Scholar]
- Subchev M. 2014. The genus Branchiobdella Odier, 1823 (Annelida, Clitellata, Branchiobdellida): a review of its european species. Acta Zool Bulg 66: 5–20. [Google Scholar]
- Subchev MA, Rimcheska BJ. 2021. Description of Branchiobdella bulgariensis sp. n. (Branchiobdellida) from Bulgaria. Acta Zool Bulg 73: 331–338. [Google Scholar]
- Subchev MA, Vaitonis G, Višinskienė G, Rimcheska BJ, Vagalinski BL. 2024. The genus Branchiobdella Odier, 1823 (Annelida: Branchiobdellida) in Lithuania, with an overview and an identification key to the species in the Baltic countries. Acta Zool Bulg 76: 3–10. [Google Scholar]
- Szenejko M, Eljasik P, Panicz R, Śmietana P. 2023. Declining populations of Astacus astacus drag European Branchiobdella parasita (Annelida: Clitellata) ectosymbionts to serious biodiversity loss. Glob Ecol Conserv 48: e02719. [Google Scholar]
- Tamura K, Stecher G, Kumar S. 2021. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol 38: 3022–3027. [CrossRef] [PubMed] [Google Scholar]
- Trifinopoulos J, Nguyen LT, von Haeseler A, Minh BQ. 2016. W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res 44: W232–W235. [Google Scholar]
- Trontelj P, Machino Y, Sket B. 2005. Phylogenetic and phylogeographic relationships in the crayfish genus Austropotamobius inferred from mitochondrial COI gene sequences. Mol Phylogenet Evol 34: 212–226. [Google Scholar]
- Villesen P. 2007. FaBox: an online toolbox for fasta sequences. Mol Ecol Notes 7: 965–968. [Google Scholar]
- Vitecek S, Graf W, Previšić A, Kučinić M, Oláh J, Bálint Keresztes L, Pauls SU, Waringer J. 2015. A hairy case: The evolution of filtering carnivorous Drusinae (Limnephilidae, Trichoptera). Mol Phylogenet Evol 93: 249–260. [Google Scholar]
- Vlach P, Šrámková L, Fialová K, Nováková K. 2017. The occurrence of branchiobdellidans on stone crayfish (Austropotamobius torrentium) in the Czech Republic. Knowl Manag Aquat Ecosyst 418: 39. [Google Scholar]
Cite this article as: Klobučar G, Vucić M, Anđelić Dmitrović B, Podnar M, Subchev MA, Maguire I. 2026. Newly discovered diversity within European branchiobdellids - a story of co-evolution and a need for conservation. Knowl. Manag. Aquat. Ecosyst., 427, 13. https://doi.org/10.1051/kmae/2026006
All Tables
Crayfish basibiont species and European branchiobdellid species. (+ confirmed findings in the literature, ? – rare findings – possible transfer from other crayfish species or misidentification, cells with grey background – species sequenced in this study).
Inter- and intraspecific divergence over sequence pairs for Branchiobdella species. Average and range values (minimum and maximum values are shown in parentheses) of uncorrected p-distances among and within studied Branchiobdella species are presented.
Intraspecific divergence over sequence pairs for groups recognized by mPTP analysis. P-distances for groups defined by using species delimitation method (NA – not applicable, only one sequence).
Interspecific divergence over sequence pairs for groups within B. hexadonta recognized by mPTP analysis. Basibionts of specific groups are indicated.
Interspecific divergence over sequence pairs for groups within B. parasita recognized by mPTP analysis. Basibionts of specific groups are indicated.
Interspecific divergence over sequence pairs for groups within B. pentadonta recognized by mPTP analysis. Basibionts of specific groups are indicated.
Interspecific divergence over sequence pairs for groups within B. italica recognized by mPTP analysis. Basibionts of specific groups are indicated.
Interspecific divergence over sequence pairs for groups within B. kozarovi recognized by mPTP analysis. Basibionts of specific groups are indicated.
All Figures
 |
Fig. 1 Locations with sequenced branchiobdellidans (B. astaci, B. balcanica, B. hexadonta, B. italica, B. kozarovi, B. parasita, and B. pentadonta). Areas of distribution for freshwater crayfish: Au. pallipes s.l., Au. torrentium, and A. astacus. A) Europe and B) South-eastern Europe. Crayfish distributions were taken from the IUCN Red List and adapted for this study: A. astacus (Edsman et al., 2010), Au. torrentium (Pârvulescu et al., 2025), and Au. pallipes (Füreder et al., 2010). Maps were produced using QGIS software v. 3.40 (QGIS, 2025). |
| In the text | |
 |
Fig. 2 The maximum-likelihood tree of genus Branchiobdella based on cytochrome c oxidase subunit I gene analysis was inferred with IQ-Tree (Nguyen et al., 2015). The numbers on the branches present ultrafast bootstrap values (≥0.7). Sequences were annotated with haplotype number according to Table S1. As presented in the legend, the symbols next to the haplotype names indicate the crayfish host. Asterisks (*) represent the new haplotypes obtained in this study. Groups recognized by mPTP analysis are displayed on the right side of the phylogram before the branchdiobdellidan species affiliation or name. |
| In the text | |
 |
Fig. 3 Time-calibrated Bayesian phylogenetic tree inferred from mitochondrial COI sequences. The divergence times are depicted as median values and 95% HPD intervals. Circles on nodes represent clade support (Bayesian posterior probabilities, PP) as shown in the figure. 15 groups revealed by species delimitation are shown as well as employed haplotypes (details are provided in Tab. S1). Further, the names of branchiobdellid species are provided next to branches. Nodes A and B represent calibration points: A – the split of North American and European Astacids, and B – the split of Au. torrentium and Au. pallipes s.l. |
| In the text | |
 |
Fig. 4 Distribution of three well-supported groups (9, 10, and 11) within B. hexadonta. |
| In the text | |
 |
Fig. 5 Distribution of three well-supported groups (13, 14, and 15) within B. parasita. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


