| Issue |
Knowl. Manag. Aquat. Ecosyst.
Number 417, 2016
Topical issue on Crayfish
|
|
|---|---|---|
| Article Number | 1 | |
| Number of page(s) | 8 | |
| DOI | https://doi.org/10.1051/kmae/2015037 | |
| Published online | 18 January 2016 | |
Monitoring of white-clawed crayfish (Austropotamobius pallipes) population during a crayfish plague outbreak followed by rescue
Suivi d’une population d’écrevisse à pattes blanches Austropotamobius pallipes durant une mortalité massive dûe à la peste et son opération de sauvetage
1
Office National de l’Eau et des Milieux Aquatiques ONEMA
Délégation régionale Alsace, Lorraine, Champagne Ardenne, 23 rue des Garennes, 57155
Marly,
France
2
Laboratoire Ecologie et Biologie des Interactions – UMR CNRS 7267
Equipe Ecologie Evolution Symbiose, Bâtiment B8-B35, 6, rue Michel Brunet, TSA 51106, 86073
Poitiers Cedex,
France
3
115 Weirview Drive,
Stillorgan, Co.
Dublin, Ireland
⋆ Corresponding author:
This email address is being protected from spambots. You need JavaScript enabled to view it.
Received:
15
July
2015
Revised:
11
November
2015
Accepted:
14
November
2015
Abstract
A mass mortality was detected in the downstream section of one of the most extensive French populations of the endangered white-clawed crayfish, Austropotamobius pallipes, on June 26, 2013. This population occupied a 12 km stretch of the La Lucelle brook, with an estimated size of around 150 000 individuals. The presence of the crayfish plague pathogen was quickly diagnosed as the cause of the mortality, and monitoring was carried out to follow the spread of the disease from 15 July 2013 for one year. Results showed that after a fast spread between 15 and 25 July 2013 (upstream progression of mortality for about 4 km), the mortality front was limited to a stretch of a few hundred meters from August until December 2013. During winter, mortality was always observed, confirming that disease remained active. In April 2014, the mortality front was halted by a large dam in the brook (2 m high), 0.56 km from brook source. Two months later, 30 live crayfish were observed above the dam. On the 30 August 2014, no crayfish were found above the dam. Infected individuals analysed for microsatellite markers confirmed the Pacifastacus leniusculus strain of Aphanomyces astaci at the origin of this outbreak. Before the crayfish plague spread upstream of the large dam, a sample of 576 individuals was collected from upstream of the dam and translocated to another stream in the same French department. In July 2014, observations by night confirmed the presence of translocated white-clawed crayfish in the receiving brook.
Résumé
Une mortalité massive a été détectée dans la section aval de l’une des plus importantes populations françaises d’écrevisse à pattes blanches, Austropotamobius pallipes le 16 juin 2013. Cette population occupait 12 km du cours d’eau de la Lucelle, avec une taille de population estimée à 150 000 individus. La présence du pathogène responsable de la peste de l’écrevisse a été rapidement diagnostiquée comme la cause de cette mortalité, et un suivi a été réalisé à partir du 15 juillet 2013 pendant un an. Les résultats ont montré qu’après une rapide progression de la mortalité entre le 15 et 25 juillet 2013 (de 4 km vers l’amont), le front de mortalité a ensuite stagné sur quelques centaines de mètres d’août à décembre 2013. Durant l’hiver, la pathologie est restée active. En avril 2014, la mortalité a atteint un conséquent barrage (2 m de haut) situé à 0,56 km de la source. Deux mois plus tard, le 30 juin, 30 écrevisses vivantes ont été observées en amont du barrage. Le 30 août, aucun individu n’a été retrouvé en amont de celui-ci. L’analyse moléculaire par microsatellite des individus infectés confirme que la souche d’Aphanomyces astaci incriminée est celle de Pacifastacus leniusculus. Le 22 avril, le statut d’infection des individus situés à 40 m en amont du barrage a confirmé l’absence d’infection de ceux-ci. Une translocation de 576 individus non infectés collectés en amont du barrage, a été effectuée dans un cours d’eau situé dans le même département. En juillet 2014, des observations de nuit à la lampe ont confirmé la présence d’écrevisses dans le cours d’eau receveur.
Key words: Disease outbreak / monitoring / molecular detection / management / translocation
Mots clés : Mortalité massive / monitoring / détection moléculaire / gestion / translocation
© M. Collas et al., published by EDP Sciences, 2016
 This is an Open Access article distributed under the terms of the Creative
Commons Attribution License CC-BY-ND (http://creativecommons.org/licenses/by-nd/4.0/), which permits
unrestricted use, distribution, and reproduction in any medium, provided the original
work is properly cited. If you remix, transform, or build upon the material, you may not
distribute the modified material.
This is an Open Access article distributed under the terms of the Creative
Commons Attribution License CC-BY-ND (http://creativecommons.org/licenses/by-nd/4.0/), which permits
unrestricted use, distribution, and reproduction in any medium, provided the original
work is properly cited. If you remix, transform, or build upon the material, you may not
distribute the modified material.
Research Article
1 Introduction
Over recent decades, the distribution of the white-clawed crayfish Austropotamobius pallipes (Lereboullet, 1838) has become restricted and fragmented throughout its natural range in Europe (Souty-Grosset et al., 2006; Holdich et al., 2009; Kouba et al., 2014). In France, the situation is particularly alarming. A survey conducted by the National Office for Water and Aquatic Habitats (ONEMA) has shown that the number of A. pallipes populations has considerably declined over the last 30 years (Collas et al., 2007). In a detailed case study in the Poitou-Charentes region of France, Bramard et al. (2006) reported a 70% diminution of the number of populations between 1978 and 2006, from 148 populations to 45, with a further loss of 40% in the last 6 years. As a result of this decline, A. pallipes is now listed in the IUCN Red Book of endangered species since 2010 (Füreder et al., 2010) also under Annex 2 of the EU Habitats Directive as a species requiring special conservation measures.
Among the factors given to explain the disappearance of most A. pallipes populations, the crayfish plague caused by the oomycete pathogen Aphanomyces astaci Schikora, 1906 is one of the main reasons, having affected many populations throughout Europe (e.g. Holdich et al., 2009; Alonso et al., 2000; OIE, 2009). The first European outbreak of the plague was recorded in 1859 in northern Italy (Alderman, 1996). In France, the disease first appeared in 1874 at Plateau des Langres (Alderman, 1996) and is considered to be the second wave of mass mortalities in Europe. Since then, large outbreaks due to the plague have been reported in many European countries, most of them associated with the presence of American crayfish species acting as potential carriers of A. astaci (e.g., Filipová et al., 2013). Several American crayfish species have been successfully introduced into Europe, including mainly the spiny-cheek crayfish Orconectes limosus (Rafinesque, 1817), the signal crayfish Pacifastacus leniusculus (Dana, 1852) and the red swamp crayfish Procambarus clarkii (Girard, 1852). As they are invasive, they are now very common in much of Europe (Kouba et al., 2014). Recently, several molecular methodologies have been developed to detect A. astaci directly from infected crayfish tissues (Oidtmann et al., 2006; Vrålstad et al., 2009, 2014; Tuffs and Oidtmann, 2011; Grandjean et al., 2014) and have improved the possibilities for screening for this pathogen (Kozubíková et al., 2008; 2011b; Aquiloni et al., 2010; Vrålstad et al., 2011; Pârvulescu et al., 2012). These studies confirmed that all of these North American crayfish species can carry crayfish plague and so can freshwater crab species (Svoboda et al., 2014; Schrimpf et al., 2014). In a large scale epidemiology screening in France, Filipova et al. (2013) reported that 29 out of 54 populations of P. leniusculuswere infected by A. astaci.
Five groups of A. astaci strains have now been described based on RAPD analysis on pure culture: group A that comprises A. astaci strains isolated from infected European crayfish species (Astacus astacus (Linnaeus, 1758) and Astacus leptodactylus (Eschscholtz, 1823)) and with an original host still remaining unknown; groups B and C that contain strains isolated from P. leniusculus of Californian and Canadian origin, respectively (Huang et al., 1994); group D, isolated from P. clarkii and showing physiological adaptations to warm waters (Diéguez-Uribeondo et al., 1995); and group E isolated from O. limosus (Kozubíková et al., 2011a). These genotype groups have been recently confirmed using specific microsatellite markers from A. astaci (Grandjean et al., 2014). In a comparative study to test the virulence among strains, Makkonen et al. (2012) reported that strains from group B are more virulent than strains from group A.
When plague occurs in a European crayfish population, the population is generally eliminated within short periods of time (Kozubíková et al., 2008; OIE, 2009) although a few cases of chronic infections have been recently reported for indigenous European crayfish species (Caprioli et al., 2013; Kušar et al., 2013; Viljamaa-Dirks et al., 2013). In some cases, the presence of a pond or dam may limit the upstream spread of disease (Kozubíková-Balcarova et al., 2014). Due to its virulence and devastating impact on indigenous European crayfish species, the crayfish plague pathogen has been classed among the world’s 100 worst invasive alien species (Lowe et al., 2004).
However, although mass mortalities due to the plague are widespread throughout Europe, generally no detailed monitoring has been performed to estimate the spread of the disease over time. Among the symptoms that could be observed during a crayfish plague outbreak, is the presence of moribund crayfish with behavioral abnormalities, such as the phenomenon of “walking on stilts” in which infected individuals stand on the tips of their walking legs (Alderman and Polglase, 1988) or a slow or no tail escape response. This behavior affects all age classes of the population. Most outbreaks are indirectly linked to the presence of American crayfish populations living close to a native crayfish population (Viljamaa-Dirks et al., 2013) although in some cases, the link is not so straightforward (Kozubíková-Balcarova et al., 2014).
In June 2013, a massive outbreak episode affected one of the largest and most widespread populations of white-clawed crayfish in a border brook between France and Switzerland. This population extended over 12 km and was considered as a potential reservoir for restocking (Reichen and Periat, 2002; Stucki and Zaugg, 2006) The presence of moribund and dead crayfish was observed by Swiss authorities on 21 June 2013 in a downstream part. Plague was suspected and quickly confirmed by molecular analysis in the beginning of July 2013. This outbreak gave us the opportunity to follow the spread of mortality front for one year. The A. astaci strain involved in this outbreak was characterized using specific molecular makers recently developed by Grandjean et al. (2014). Finally, crayfish from the upstream part, free of plague at the time of translocation, were rescued by French authorities of ONEMA and translocated to a suitable brook free of non-indigenous crayfish.
 |
Fig. 1 Photo of dead crayfish found in sector A,17 July 2014. |
2 Materials and methods
La Lucelle (Long. 7°15′51′′E, Lat. 47°25′34′′N) is a 27 km long brook, on the border between France and Switzerland (Figure 1), and it is located in the Rhine drainage. The headwaters are dammed at 0.5 km from the source, forming a lake of 5 ha. This brook runs through forested calcareous hillsides and flows into the Birse River.
Monitoring dates of crayfish plague spread in the Lucelle. Observations were made by day or by light during the night.
The brook varies from 10 to 80 cm in depth and 4 to 6 m in width. Flow rates and sediment granulometry are also varied. Stones and gravel (80%) are predominant whereas sand and clay are less well represented. There is intact riparian vegetation with the presence of roots of ash and alder trees. The crayfish population occurs over 12 km in the most upstream part of the brook. Population size was estimated by CRM (capture-mark-recapture) in 2009 by ONEMA.
2.1 Monitoring and sample collections
The outbreak was initially reported on 21 June 2013 by the Swiss fishery authorities in the most downstream part of the population. French authorities of ONEMA were informed on 4 July 2013 and fourteen field surveys were conducted from 15 July 2013 until 10 July 2014 (Table 1).
Field operations were carried out by day or by night, using lights. For each field survey, 6 people divided into 3 teams walked along the banks to detect dead, dying or living specimen in order to delimit the mortality front. In daylight, hand-searches under stones were also performed in downstream direction (Table 1). Additionally, traps baited with sardines were used on 22 April to confirm the mortality front and to collect samples to analyze the status of infection of individuals below the dam (Table 1).
The numbers of crayfish observed live or dead were recorded on 17 July and 12 August 2013, and 17 March 2014 (Table 1).
2.2 Detection of Aphanomyces astaci by quantitative real time-PCR
Sampled crayfish were stored in 96% ethanol. Tissue from one half of the soft abdominal cuticle and one uropod was dissected from each crayfish using sterile instruments. Dissected tissues from each individual were placed in a single 1.5 mL tube, dried and stored in a deep-freezer at −80 °C. Before further processing, 360 μL of ATL Buffer from the DNeasy tissue kit (Qiagen) and 40 μL of proteinase K solution were added to the dissected material. The mixture was then crushed by one scoop (ca. 50 μL) of stainless steel beads (1.6 mm diameter) using a BBX24B Bullet Blender (Next Advance) for 10 min at maximum speed. 400 μL of the Buffer AL were added to DNA extractions from the crushed cuticle, then followed the rest of the spin-column protocol of the DNeasy tissue kit.
Isolated material was then tested for the presence of A. astaci by the quantitative TaqMan® MGB real-time PCR developed by Vrålstad et al. (2009), using the LightCycler 480 Instrument (Roche). Experimental procedures were identical to those used by Filipova et al. (2013). Based on their PFU values, samples were classified into semi-quantitative categories of pathogen load, ranging from A0 (no traces of A. astaci DNA) to A7 (extremely high amount of A. astaci DNA in the sample), as proposed by Vrålstad et al. (2009).
2.2.1 (i) Diagnosis of plague detection and strain determination
10 dead individuals of A. pallipes were sampled on 15 July 2013 for plague detection and genotyping. Eight microsatellite loci developed by Grandjean et al. (2014) were genotyped on 5 samples. All of them were positive for A. astaci presence, with level of infection ranging from A4 to A6 (moderate to high infection levels) according to Vrålstad et al. (2009). All experimental procedures for microsatellite genotyping were identical to those described in Grandjean et al. (2014).
2.2.2 (ii) Detection of plague-free crayfish for possible translocation
On 18 December 2013, 15 crayfish from small brook sections downstream and upstream of the dam ‘f’ (Figure 2) were analysed using the same protocol to test if there are some plague-free crayfish for translocation purposes.
2.2.3 (iii) Characterization of infected crayfish along mortality front
On 21 April 2014, 9 baited traps were placed from the mortality front delimited on 3 April located at 200 m downstream of the dam to check the infection status of collected crayfish. Seven traps were placed downstream of the dam (at 200 m, 110 m, 80 m, 60 m, 40 m, 20 m, 10 m) and two upstream of the dam (10 m, 130 m) (Figure 2).
3 Results
3.1 Survey
The first dead crayfish were observed on 21 June 2013 in the most downstream part of the crayfish population. On 15 July 2013, 3 sectors were defined. Sector A (coloured red in Figure 2) had mostly dead crayfish exhibiting a white mycelium on the body (Figure 1). In sector B (coloured yellow in Figure 2) mortality was in progress, and crayfish were outside their shelters, lacked any escape reaction when we tried to catch them. Some crayfish lay on their backs, slowly moving their legs. In this sector, no crayfish covered by mycelium were observed. Sector C (coloured blue in Figure 2) was the most upstream sector, and presumably free of the infection; all crayfish were in their shelters and showed normal escape behaviour. Mortality front was defined as a border between sector A and sector B while disease front was situated between sector B and sector C. At this date the disease front was located between km 5 and 6 (Figure 2a, Table 1).
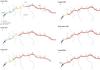 |
Fig. 2 Monitoring of outbreak from 15 July to 30 June 2014. Numbers refer to the official Km metrics from the road along the road. The black line upstream represents the dam. No mortality detected (sector A, in blue), mortalities in progress (sector B, in yellow), mass mortality (sector C, in red). |
Surveys every two days showed that the disease front was progressing upstream of 3 km between the 15 july to 25 July 2013. Ten days later, the spread of disease was limited to 300 m further upstream (6 August 2013) (Table 1). Then, no progression of the disease was reported until the end of December 2013. Two and half months later, the mortality front was located 520 m upstream, a spread of around 170 m per month, just 300 m downstream of the dam (Figures 2 and 3, Table 1). Live crayfish with normal behaviour were observed 4 m upstream from the mortality front. At the end of April 2014, dead crayfish were observed dowstream of the dam. Two months later (30 June 2014), 30 live crayfish were observed above the dam. On the 30 August 2014, no crayfish were found above the dam. The population is considered to have been lost to crayfish plague.
 |
Fig. 3 Photo of 2 m high dam with outfall pipes in the bottom located in upstream section. |
3.2 Molecular diagnostics
Genotyping of the A. astaci strain from dead individuals collected on 15 July 2013 showed allele size matching those found for genotype group B (strain originated from P. leniusculus).
Infection rate of crayfish sampled along the mortality front and up to the dam on 22 April 2014.
Disease levels detected in specimens are summarised in Table 2. The levels of infection for crayfish in the mortality sector ranged from A3 to A6 (Table 2). The number of crayfish collected by traps on 22 April 2014 and their infection status are reported in Table 2.
The analysis of infection status of crayfish sampled by traps on 22 April 2014 showed that live crayfish collected under stones, located at 40 m above the mortality front, were all infected. At this period, samples from above the dam were free of plague.
3.3 Translocation
On 3–4 April 2014 technical staff from ONEMA rescued 572 crayfish (307 males, 249 females, 15 juveniles) from above the dam, and moved them to the Lutter Brook (Long. 7°22′53′′E, Lat. 47°28′01′′N) located in the same French department. Its physical morphology is closely similar to that at sites with the white-clawed crayfish in this region, with stones and rocks as the dominant substrates (90% of surface) (Collas, pers. com.). On 29 June and 5 September 2014, 3 and 2 crayfish respectively were observed during the night search in the receiving brook.
4 Discussion
This study describes a rapid mass mortality due to crayfish plague in Europe in one of the densest French populations of white-clawed crayfish. The mortality affected all size ranges of crayfish over the 12 km of brook inhabited by A. pallipes. It started around 21 June 2013. The population was considered extinct at the end of August 2014. As generally observed in plague outbreaks, the disease first occurred downstream and spread quickly upstream.
However, the speed of spread over the survey period was quite variable. From the start of the outbreak in mid-June to the beginning of August, an extremely rapid spread of infection was reported, with the disease front estimated to move around 3 km upstream. Then, a stagnation of the disease front was observed from August to the end of 2013. During winter, the disease front spread 520 m upstream. This result showed that the outbreak was active even during winter, an unexpected finding according to the literature (OIE, 2009) but also reported by Kozubíková-Balcarova et al. (2014) in some outbreaks in the Czech Republic.
These differences in rate of spread of the infection in this population were difficult to explain, but several hypotheses could be drawn. (i) It is known that crayfish density can act on the quantity of spore production during an outbreak (Caprioli et al., 2013). If so, we could expect more spore production in areas of high density and a quick spread of infection. Conversely, where densities are lower, fewer zoospores are produced, which may limit the speed of infection. Over the 12 km colonized by crayfish, density may have fluctuated along the brook. (ii) Water temperature is also known to have an impact on the production of zoospores (Strand et al., 2012). At low temperatures the life cycle of A. astaci is longer, limiting the speed of the disease because the production of zoospores was reduced. In our case, it could explain the limited progression of the outbreak during the winter period, but not the stabilization of the disease front in August and September, when water temperature was highest. (iii) Natural waterfalls present between km 3 and 4 could act as a barrier limiting the upstream progression of disease.
One year after the beginning of the outbreak, crayfish mortality had affected the whole population. In a case study on status and recovery of autochthonous crayfish populations after plague outbreaks, Kozubíková-Balcarova et al. (2014) reported that the presence of migration barriers (1 or 2 m high) on the streams stopped the spread of the disease in three cases. Migration barriers as effective measures to prevent upstream spread of the disease had also been reported by Rahel (2013). More relevant examples of functional migration barriers were also reported in Vrålstad et al. (2011). In our study, the 2 m high dam did not prevent the upstream spread of the disease. However, as the dam has an outfall pipe at the bottom, crayfish or fish could be able to pass through when the flow is not too strong. Other vectors such as other animals (e.g. semiaquatic mammals or birds) may have carried spores from down to and above the dam.
Due to the extreme rapidity of the spread of the disease and the total mortality of infected crayfish, the characterization of strain B involved in this outbreak was not surprising. This strain is considered highly virulent and has been involved in massive outbreaks, particularly in Norway, Finland, Czech Republic and France (Viljamaa-Dirks et al., 2013; Kozubíková-Balcarova et al., 2014; Grandjean et al., 2014; Strand et al., 2014). This strain is specific to P. leniusculus which often shares the habitat of the white-clawed crayfish in France (Bramard et al., 2006; Filipova et al., 2013) which means that this strain is particularly well adapted to the water conditions of the La Lucelle brook. No signal crayfish were observed during the survey but several populations are established in the same hydrographic basin. Underlying this study is the persistent question of how the disease arrived to infect the La Lucelle population. Unfortunately, we do not have a clear answer. The spores of the fungus can be carried by a variety of means: in water, on fish, in mud on boots, and on fishing equipment (OIE, 2009; Reynolds, 1988). The non-observation of signal crayfish was not a guarantee of its absence. Indeed, signal crayfish could go undetected for decades after introduction in large aquatic ecosystems such as lakes or large rivers (Johnsen et al., 2007; Vrålstad et al., 2011). The presence of signal crayfish could be due to a natural colonization from specimens of populations already established in the same drainage or by illegal introduction by man. Introduction by man is by far the most common scenario of signal crayfish spread in France (Bramard et al., 2006). Further monitoring using cages with indigenous crayfish and/or water monitoring of A. astaci spore content (Strand et al., 2014) in the years to come can reveal if the infection has disappeared (expected in case of no crayfish) or remain (expected if signal crayfish are present). In the case of plague-free situation, this brook could be restocked in the next few years. It would be interesting to check the infection status of signal populations around and inform anglers about the risk of disease propagation. The disease is very difficult to control and outbreaks are impossible to predict. However it is very easy to prevent by simple biosecurity measures (cleaning and drying gear).
The analysis of infection status of individuals caught by traps from the mortality front up to the dam shows that crayfish were infected 100 m above the mortality front. Probably this distance might have been longer if there was no dam. Then, a rescue attempt of live crayfish above the mortality front would be very risky. In our case, the presence of a dam upstream combined with negative results from molecular plague detection offered us the possibility to rescue some crayfish. The precautionary principle on target habitats (Souty-Grosset and Reynolds, 2009) was applied in introducing the rescued crayfish into a suitable brook free of crayfish and located in the same hydrographic basin.
References
- Alderman D.J., 1996. Geographical spread of bacterial and fungal diseases of crustaceans. Revue Scientifique et Technique – Office International des Epizooties, 15, 603–632. [Google Scholar]
- Alderman D.J. and Polglase J.L., 1988. Pathogens, parasites and commensals. In: Holdich D.M. and Lowery R.S. (eds.), Freshwater Crayfish: Biology, Management and Exploitation. CroomHeim, London, pp. 167–212. [Google Scholar]
- Alonso F., Temino C. and Diéguez-Uribeondo J., 2000. Status of the white-clawed crayfish, Austropotamobius pallipes (Lereboullet, 1858), in Spain: distribution and legislation. Bulletin Français de la Pêche et de la Pisciculture, 356, 31–54. [Google Scholar]
- Aquiloni L., Tricarico E. and Gherardi F., 2010. Crayfish in Italy: distribution, threats and management. Int. Aquat. Res., 2, 1–14. [Google Scholar]
- Bramard M., Demers A., Trouilhé M.C., Bachelier E., Dumas J.C., Fournier C., Broussard E., Robin O., Souty-Grosset C. and Grandjean F., 2006. Distribution of indigenous and non-indigenous crayfish populations in the Poitou-Charentes Region (France): Evolution over the past 25 years. Bull. Fr. Pêche Piscic., 380–381, 857–865. [CrossRef] [EDP Sciences] [Google Scholar]
- Caprioli R., Cargini D., Marcacci M., Camm C., Giansante C. and Ferri, N., 2013. Self-limiting outbreak of crayfish plague in an Austropotamobius pallipes population of a river basin in the Abruzzi region (central Italy). Dis. Aquat. Organ., 103, 149–156. [CrossRef] [PubMed] [Google Scholar]
- Collas M., Julien C. and Monnier D., 2007. La situation des écrevisses en France. Résultats des enquêtes nationales réalisées entre 1977 et 2006 par le Conseil Supérieur de la pêche. Bull. Fr. Pêche Piscic., 386, 1–38. [Google Scholar]
- Diéguez-Uribeondo J., Huang T.S., Cerenius L. and Soderhall K., 1995. Physiological adaptation of an Aphanomyces astaci strain isolated from the freshwater crayfish Procambarus clarkii. Mycological Res., 99, 574–578. [CrossRef] [Google Scholar]
- Filipová L., Petrusek A., Matasová K., Delaunay C. and Grandjean F., 2013. Prevalence of the Crayfish Plague pathogen Aphanomyces astaci in populations of the signal crayfish Pacifastacus leniusculus in France: Evaluating the threat to native crayfish. PLoS One, 8, e70157. Doi:10.1371/journal.pone.0070157. [Google Scholar]
- Füreder L., Gherardi F., Holdich D., Reynolds J., Sibley P. and Souty-Grosset C., 2010. Austropotamobius pallipes. In: IUCN 2011. IUCN Red List of Threatened Species. Version 2014.1. http://www.iucnredlist.org/. [Google Scholar]
- Grandjean F., Vrålstad T., Diéguez-Uribeondo J., Jeli M., Mangombi J., Delaunay C., Filipová L., Rezinciuc S., Kozubíkov-Balcarová E., Guyonnet D., Viljamaa-Dirks S. and Petrusek A., 2014. Microsatellite markers for direct genotyping of the crayfish plague pathogen Aphanomyces astaci (Oomycetes) from infected host tissues. Veter. Microbiol., 170, 317–324. [CrossRef] [Google Scholar]
- Holdich D.M., Reynolds J., Souty-Grosset C. and Sibley P.J., 2009. A review of the ever increasing threat to European crayfish from non-indigenous crayfish species. Knowl. Manag. Aquat. Ecosyst., 394–395, 1–46. [Google Scholar]
- Huang T.S., Cerenius L. and Soderhall K., 1994. Analysis of genetic, diversity in crayfish plaguefungus, Aphanomyces astaci, by random amplification of polymorphic DNA. Aquaculture, 126, 1–10. [CrossRef] [Google Scholar]
- Johnsen S.I, Taugbol T., Andersen O., Mirseth J. and Vrålstad T., 2007. The first record of non-indigenous signal crayfish Pacifastacus leniusculus in Norway. Biological Invasions, 9, 939–941. [CrossRef] [Google Scholar]
- Kouba A., Petrusek, A. and Kozák P., 2014. Continental-wide distribution of crayfish species in Europe: update and maps. Knowl. Manag. Aquat. Ecosyst., 413, 1–31. [Google Scholar]
- Kozubíková E., Petrusek A., Uriš Z., Martín M.P., Diéguez-Uribeondo J. and Oidtmann B., 2008. The old menace is back: recent crayfish plague outbreaks in the Czech Republic. Aquaculture, 274, 208–217. [CrossRef] [Google Scholar]
- Kozubíková E., Viljamaa-Dirks S., Heinikainen S. and Petrusek A., 2011a. Spiny-cheek crayfish Orconectes limosus carry a novel genotype of the crayfish plague pathogen Aphanomyces astaci. J. Invertebr. Pathol., 108, 214–216. [CrossRef] [PubMed] [Google Scholar]
- Kozubíková E., Vrålstad T., Filipová L. and Petrusek A., 2011b. Re-examination of the prevalence of Aphanomyces astaci in North American crayfish populations in Central Europe by TaqMan® MGB real-time PCR. Dis. Aquat. Organ., 97, 113–125. [CrossRef] [PubMed] [Google Scholar]
- Kozubíková-Baraclová E., Beran L., Duris Z., Fischer D., Horka I., Svobodova J. and Petrusek A., 2014. Status and recovery of indigenous crayfish populations after recent crayfish plague outbreaks in the Czech Republic. Ethol. Ecol. Evol., 26, 299–319. [CrossRef] [Google Scholar]
- Kušar D., Vrezec A., Ocepek M. and Jeničič V., 2013. Aphanomyces astaci in wild crayfish populations in Slovenia: first report of persistent infection in a stone crayfish Austropotamobius torrentium population. Dis. Aquat. Organ., 103, 157–169. [Google Scholar]
- Lowe S., Browne M., Boudjelas S. and De Poorter M., 2004. 100 of the world’s worst invasive alien species.A selection from the Global Invasive Species Database. The Invasive Species Specialist Group (ISSG), a specialist group of the Species Survival Commission (SSC) of the IUCN, Gland, Switzerland. [Google Scholar]
- Makkonen J., Jussila J., Kortet R., Vainikka A. and Kokko H., 2012. Differing virulence of Aphanomyces astaci isolates and elevated resistance of noble crayfish Astacus astacus against crayfish plague. Dis. Aquat. Organ., 102, 129–136. [Google Scholar]
- Oidtmann B., Geiger S., Steinbauer P., Culas A. and Hoffmann R.W., 2006. Detection of Aphanomyces astaci in North American crayfish by polymerase chain reaction. Dis. Aquat. Organ., 72, 53–64. [CrossRef] [PubMed] [Google Scholar]
- OIE (Office International des Epizooties), 2009. Crayfish plague (Aphanomyces astaci). In: Manual of diagnostic tests for aquatic animals, 6th edition. Office international des épizooties, Paris, Chap. 2.2.1, pp. 63–77. http://www.oie.int. [07 July 2014]. [Google Scholar]
- Pârvulescu L., Schrimpf A., Kozubíková E., Resino S.C., Vrålstad T., Petrusek A. and Schulz R., 2012. Invasive crayfish and crayfish plague on the move: first detection of the plague agent Aphanomyces astaci in the Romanian Danube. Dis. Aquat. Organ., 98, 85–94. [Google Scholar]
- Rahel F.J., 2013. Intentional fragmentation as a management strategy in aquatic systems. Bioscience, 63, 362–372. [CrossRef] [Google Scholar]
- Reichen D. and Periat G., 2002. Cartographie des Ecrevisses dans les cours d’eau du Canton du Jura. OEPN, St-Ursanne. [Google Scholar]
- Reynolds J.D., 1988. Crayfish extinctions and crayfish plague in Central Ireland. Biol. Conserv., 45, 279–285. [CrossRef] [Google Scholar]
- Schrimpf A., Schmidt T. and Schulz R., 2014. Invasive Chinese mitten crab (Eriocheir sinensis) transmits crayfish plague pathogen (Aphanomyces astaci). Aquatic Invasions, 9, 203–209. [Google Scholar]
- Souty-Grosset C. and Reynolds J.D., 2009. Current ideas on methodological approaches on European crayfish conservation and restocking procedures. Knowl. Manag. Aquat. Ecosyst., 394–395, 01. [Google Scholar]
- Souty-Grosset C., Holdich D.M., Noël P.Y., Reynolds J.D. and Haffner P., 2006. Atlas of Crayfish in Europe. Muséum national d’Histoire naturelle. Patrimoines naturels, 64, Paris. [Google Scholar]
- Strand D., Jussila J., Viljamaa-Dirks S., Kokko H., Makkonen J., Holst-Jensen A., Viljugrein H. and Vrålstad T., 2012. Monitoring the spore dynamics of Aphanomyces astaci in the ambient water of latent carrier crayfish. Veter. Microbiol., 160, 99–107. [CrossRef] [Google Scholar]
- Strand D., Jussila J., Johnsen S.I., Viljamaa-Dirks S., Edsman L., Wiik-Nielsen J., Viljugrein H., Engdahl F. and Vrålstad T., 2014. Detection of crayfish plague spores in large freshwater systems. J. Appl. Ecol., 51, 544–553. [CrossRef] [Google Scholar]
- Stucki P. and Zaugg B., 2006. Plan d’action national pour les écrevisses. OFEFP, 41 p. [Google Scholar]
- Svoboda J., Strand D.A., Vrålstad T., Grandjean F., Edsman L., Kozák P., Kouba A., Fristad R.F., Bahadir Koca S. and Petrusek A., 2014. The crayfish plague pathogen can infect freshwater-inhabiting crabs. Freshw. Biol., 59, 918–929. [CrossRef] [Google Scholar]
- Tuffs S. and Oidtmann B., 2011. A comparative study of molecular diagnostic methods designed to detect the crayfish plague pathogen, Aphanomyces astaci. Veter. Microbiol., 153, 343–353. [CrossRef] [Google Scholar]
- Viljamaa-Dirks S., Heinikainen S., Torssonen H., Pursiainen M., Mattila, J. and Pelkonen S., 2013. Distribution and epidemiology of genotypes of the crayfish plague agent Aphanomyces astaci from noble crayfish Astacus astacus in Finland. Dis. Aquat. Organ., 103, 199–208. [CrossRef] [PubMed] [Google Scholar]
- Vrålstad T., Knutsen A.K., Tengs T. and Holst-Jensen A., 2009. A quantitative TaqMan MGB real-time polymerase chain reaction based assay for detection of the causative agent of crayfish plague Aphanomyces astaci. Vet. Microbiol., 137, 146–155. [Google Scholar]
- Vrålstad T., Johnsen S.I., Fristad R.F., Edsman L. and Strand D., 2011. Potent infection reservoir of crayfish plague now permanently established in Norway. Dis. Aquat. Organ., 97, 75–83. [CrossRef] [PubMed] [Google Scholar]
- Vrålstad T., Strand, D. A, Grandjean F., Kvellestad A., Hastein T., Knutsen, A.-K., Taugbol T. and Skaar I., 2014. Molecular detection and genotyping of Aphanomyces astaci directly from preserved crayfish samples uncovers the Norwegian crayfish plague disease history. Vet. Microbiol., 173, 66–75. [CrossRef] [PubMed] [Google Scholar]
Cite this article as: M. Collas, T. Becking, M. Delpy, M. Pflieger, P. Bohn, J. Reynolds and F. Grandjean, 2016. Monitoring of white-clawed crayfish (Austropotamobius pallipes) population during a crayfish plague outbreak followed by rescue. Knowl. Manag. Aquat. Ecosyst., 417, 1.
All Tables
Monitoring dates of crayfish plague spread in the Lucelle. Observations were made by day or by light during the night.
Infection rate of crayfish sampled along the mortality front and up to the dam on 22 April 2014.
All Figures
 |
Fig. 1 Photo of dead crayfish found in sector A,17 July 2014. |
| In the text | |
 |
Fig. 2 Monitoring of outbreak from 15 July to 30 June 2014. Numbers refer to the official Km metrics from the road along the road. The black line upstream represents the dam. No mortality detected (sector A, in blue), mortalities in progress (sector B, in yellow), mass mortality (sector C, in red). |
| In the text | |
 |
Fig. 3 Photo of 2 m high dam with outfall pipes in the bottom located in upstream section. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


